HKUST scientists deploy big data methods to learn the fitness landscape of the HIV Envelope protein
Despite significant advances in medicine, an effective vaccine for the human immunodeficiency virus (HIV) is still not available, although recent hope has emerged through the discovery of antibodies capable of neutralizing diverse HIV strains. However, HIV can sometimes evade known broadly neutralizing antibody responses via mutational pathways, which makes it all the more difficult to design an effective solution.
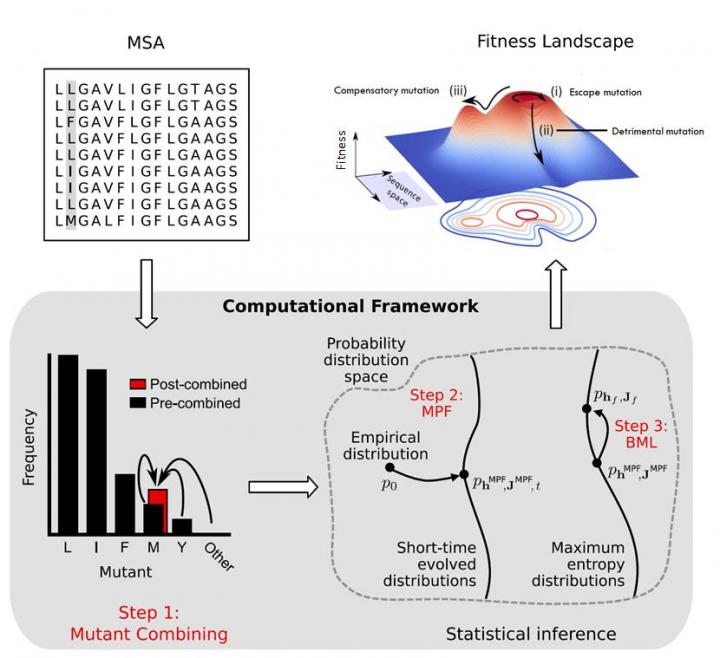
An ideal vaccine would elicit broadly neutralizing antibodies that target parts of the virus's spike proteins where mutations severely compromise the virus's fitness, or the virus' ability to reproduce and replicate. This requires knowledge of the fitness landscape, a mapping from sequence to fitness. To achieve this goal, data scientists from the HKUST have employed a supercomputational approach to estimate the fitness landscape of gp160, the polyprotein that comprises HIV's spike. The inferred landscape was then validated through comparisons with diverse experimental measurements.
Their findings were published in the journal PNAS in January 2018 (doi: 10.1073/pnas.1717765115).
"Without big data machine learning methods, it is simply impossible to make such a prediction," said Raymond Louie, co-author, Junior Fellow of HKUST's Institute for Advanced Study and Research Assistant Professor in the Department of Electronic & Computer Engineering. "The number of parameters needed to be estimated came close to 4.4 million."
The data processed by the team consisted of 815 residues and 20,043 sequences from 1,918 HIV-infected individuals.
"The computational method gave us fast and accurate results," said Matthew McKay, co-author and Hari Harilela Associate Professor in the Departments of Electronic & Computer Engineering and Chemical & Biological Engineering at HKUST. "The findings can assist biologists in proposing new immunogens and vaccination protocols that seek to force the virus to mutate to unfit states in order to evade immune responses, which is likely to thwart or limit viral infection."
"While this method was developed to address the specific challenges posed by the gp160 protein, which we could not address using methods we developed to obtain the fitness landscapes of other HIV proteins, the approach is general and may be applied to other high-dimensional maximum-entropy inference problems," said Arup K. Chakraborty, co-author and Robert T. Haslam Professor in Chemical Engineering, Physics, and Chemistry at MIT's Institute for Medical Engineering & Science. "Specifically, our fitness landscape could be clinically useful in the future for the selection of combination bnAb therapy and immunogen design."
"This is a multi-disciplinary study presenting an application of data science, and big data machine learning methods in particular, for addressing a challenging problem in biology", said McKay.

 How to resolve AdBlock issue?
How to resolve AdBlock issue?