Scientists have estimated that the age of an individual does not indicate how likely they are to be infected by SARS-CoV-2. However, the development of symptoms, the progression of the disease, and mortality are age-dependent.
There have been a large number of deaths due to the ongoing COVID-19 pandemic, and it has been shown that elderly individuals disproportionately develop severe symptoms and show higher mortality.
A team of scientists, including Associate Professor Ryosuke Omori from the Research Center for Zoonoses Control at Hokkaido University, have modeled available data from Japan, Spain, and Italy to show that susceptibility to COVID-19 is independent of age, while the occurrence of symptomatic COVID-19, severity, and mortality is likely dependent on age. Their results were published in an academic journal on October 6, 2020. 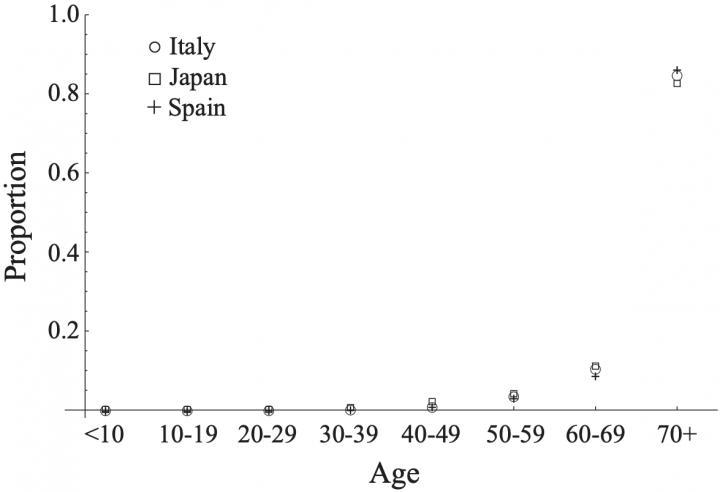 {module INSIDE STORY}
{module INSIDE STORY}
Causes of mortality in elderly individuals may be due to two factors: how likely they are to be infected due to their advanced age (age-dependent susceptibility), which is reflected in the number of cases; and, how likely they will be affected by a severe form of the disease due to their advanced age (age-dependent severity), which is reflected in the mortality rate. These factors are not fully understood for COVID-19.
The scientists chose to analyze data from Italy, Spain, and Japan to determine if any relationship between age, susceptibility, and severity. These three countries were chosen as they have well recorded, publicly available data. As of May 2020, the mortality rate (number of deaths per 100,000) was 382.3 for Italy, 507.2 for Spain, and 13.2 for Japan. However, despite the wide disparity in mortality rates, the age distribution of mortality (the proportional number of deaths per age group) was similar for these countries.
The scientists developed a mathematical model to calculate susceptibility in each age group under different conditions. They also factored in the estimated human-to-human contact level in each age group, as well as varying restriction levels for outside-home activities in the three countries.
The model showed that the susceptibility has to be unrealistically different between age groups if they assume age does not influence severity and mortality. On the other hand, the model indicated the age should not influence susceptibility but should negatively influence severity and mortality, to explain the fact that the age distribution of mortality is similar between the three countries.
Ryosuke Omori, from the Research Center for Zoonoses Control at Hokkaido University, specializes in epidemiological modeling: the use of mathematics and statistics to understand and predict the spread of diseases. Since the outbreak of COVID-19, he has turned his efforts to ascertain the true extent of the spread of the pandemic in Japan and abroad.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?