During the past years, artificial intelligence (AI) has become a key player in high-techs like drug development projects. AI tools help scientists to uncover the secret behind the big biological data using optimized computational algorithms. AI methods such as deep neural networks improve decision making in biological and chemical applications i.e., prediction of disease-associated proteins, the discovery of novel biomarkers and de novo design of small molecule drug leads. These state-of-the-art approaches help scientists to develop a potential drug more efficiently and economically. 
A research team led by Professor Hongzhe Sun from the Department of Chemistry at the University of Hong Kong (HKU), in collaboration with Professor Junwen Wang from Mayo Clinic, Arizona in the United States (a former HKU colleague), implemented a robust deep learning approach to predict disease-associated mutations of the metal-binding sites in a protein. This is the first deep learning approach for the prediction of disease-associated metal-relevant site mutations in metalloproteins, providing a new platform to tackle human diseases. The research findings were recently published in an educational journal. {module INSIDE STORY}
Metal ions play pivotal roles either structurally or functionally in the (patho)physiology of human biological systems. Metals such as zinc, iron, and copper are essential for all lives and their concentration in cells must be strictly regulated. A deficiency or an excess of these physiological metal ions can cause severe disease in humans. It was discovered that a mutation in human genome are strongly associated with different diseases. If these mutations happen in the coding region of DNA, it might disrupt metal-binding sites of the proteins and consequently initiate severe diseases in humans. Understanding of disease-associated mutations at the metal-binding sites of proteins will facilitate the discovery of new drugs.
The team first integrated omics data from different databases to build a comprehensive training dataset. By looking at the statistics from the collected data, the team found that different metals have different disease associations. A mutation in zinc-binding sites has a major role in breast, liver, kidney, immune system and prostate diseases. By contrast, the mutations in calcium- and magnesium-binding sites are associated with muscular and immune system diseases, respectively. For iron-binding sites, mutations are more associated with metabolic diseases. Furthermore, mutations of manganese- and copper-binding sites are associated with cardiovascular diseases with the latter being associated with nervous system disease as well. They used a novel approach to extract spatial features from the metal-binding sites using an energy-based affinity grid map. These spatial features have been merged with physicochemical sequential features to train the model. The final results show using the spatial features enhanced the performance of the prediction with an area under the curve (AUC) of 0.90 and an accuracy of 0.82. Given the limited advanced techniques and platforms in the field of metallomics and metalloproteins, the proposed deep learning approach offers a method to integrate the experimental data with bioinformatics analysis. The approach will help scientist to predict DNA mutations which are associated with a disease like cancer, cardiovascular diseases, and genetic disorders. 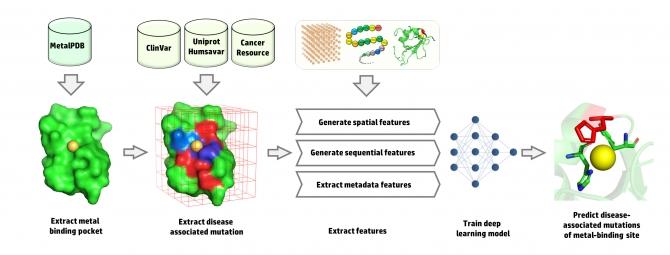 {module INSIDE STORY}
{module INSIDE STORY}
Professor Sun said: "Machine learning and AI play important roles in the current biological and chemical science. In my group, we worked on metals in biology and medicine using an integrative omics approach including metallomics and metalloproteomics, and we already produced a large amount of valuable data using in vivo/vitro experiments. We now develop an artificial intelligence approach based on deep learning to turn these raw data into valuable knowledge, leading to uncover secrets behind the diseases and to fight with them. I believe this novel deep learning approach can be used in other projects, which is undergoing in our laboratory."


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module INSIDE STORY}
{module INSIDE STORY}