Little has been known to date about how the immune system’s natural killer (NK) cells detect which cells have been infected with SARS-CoV-2. An international team of scientists led by researchers from Karolinska Institutet, ranked amongst the world's best medical schools, in Sweden now shows that NK cells respond to a certain peptide on the surface of infected cells. The study, which is published in Cell Reports, is an important piece of the puzzle in our understanding of how the immune system reacts to COVID-19.
NK cells are white blood cells that are part of the innate immune system. Unlike cells in the adaptive immune defense, they can recognize and kill cancer cells and virus-infected cells immediately without having encountered them before. This ability is controlled by a balance between the NK cells’ activating and inhibiting receptors, which can react to different molecules on the surface of other cells. 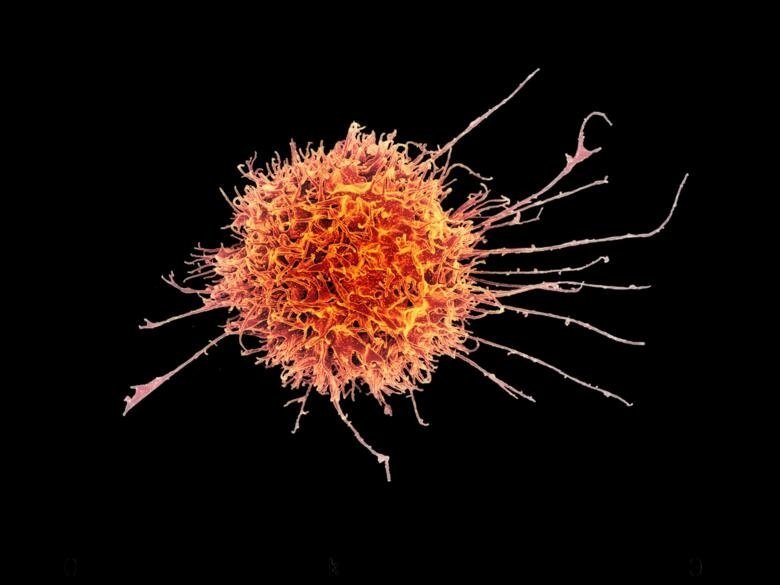
The virus is revealed by a peptide
A new study shows why certain NK cells are activated when encountering a cell infected with SARS-CoV-2. The infected cells contain a peptide from the virus that triggers a reaction in NK cells that carry a particular receptor, NKG2A, able to detect the peptide.
“Our study shows that SARS-CoV-2 contains a peptide that is displayed by molecules on the cell surface,” says Quirin Hammer, a researcher at the Center for Infectious Medicine (CIM), Karolinska Institutet. “The activation of NK cells is a complex reaction, and here the peptide blocks the inhibition of the NK cells, which allows them to be activated. This new knowledge is an important piece of the puzzle in our understanding of how our immune system reacts in the presence of this viral infection.”
The study was a major collaboration between Karolinska Institutet, Karolinska University Hospital, and research laboratories and universities in Italy, Germany, Norway, and the USA. The first phase was to test their hypothesis using supercomputer simulations that were then confirmed in the laboratory. The decisive phase was the infection of human lung cells with SARS-CoV-2 in a controlled environment, whereupon the researchers could show that NK cells with the receptor in question are activated to a greater degree than the NK cells without it.
Monitoring new virus variants
“These findings are important to our understanding of how immune cells recognize cells infected with SARS-CoV-2,” says Dr. Hammer. “This may become significant when monitoring new virus variants to determine how well the immune system responds to them.”
The study is now being followed up with the help of a biobank at Karolinska University Hospital and Karolinska Institutet containing blood samples from over 300 people treated for COVID-19 during the first wave of the pandemic.
“We’ll be examining if the composition of NK cells a person has contributes to how severe their symptoms are when infected with SARS-CoV-2,” he continues.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?