Supercomputer program advances efforts to map every neuron in worms
Researchers working towards understanding the brain in high-definition, single-cell level of detail have designed a new supercomputer program to identify each nerve cell in fluorescent microscope images of living worms. Previous attempts to automate the identification of individual nerve cells have been thwarted by the fact that the same cell can be in vastly different locations in different worms.
The worms are C. elegans, tiny roundworms common in soil and research labs around the world. Each of the 959 cells in the animals' transparent, 1 millimeter-long body has been identified, named and mapped, including their 302 nerve cells.
Scientists completed the first map of the C. elegans nervous system in 1986 and have been improving it ever since. More recent projects include OpenWorm, an ongoing global effort to design a cell-by-cell and behaviorally accurate virtual C. elegans - a research-worthy version of a Tamagotchi pet. 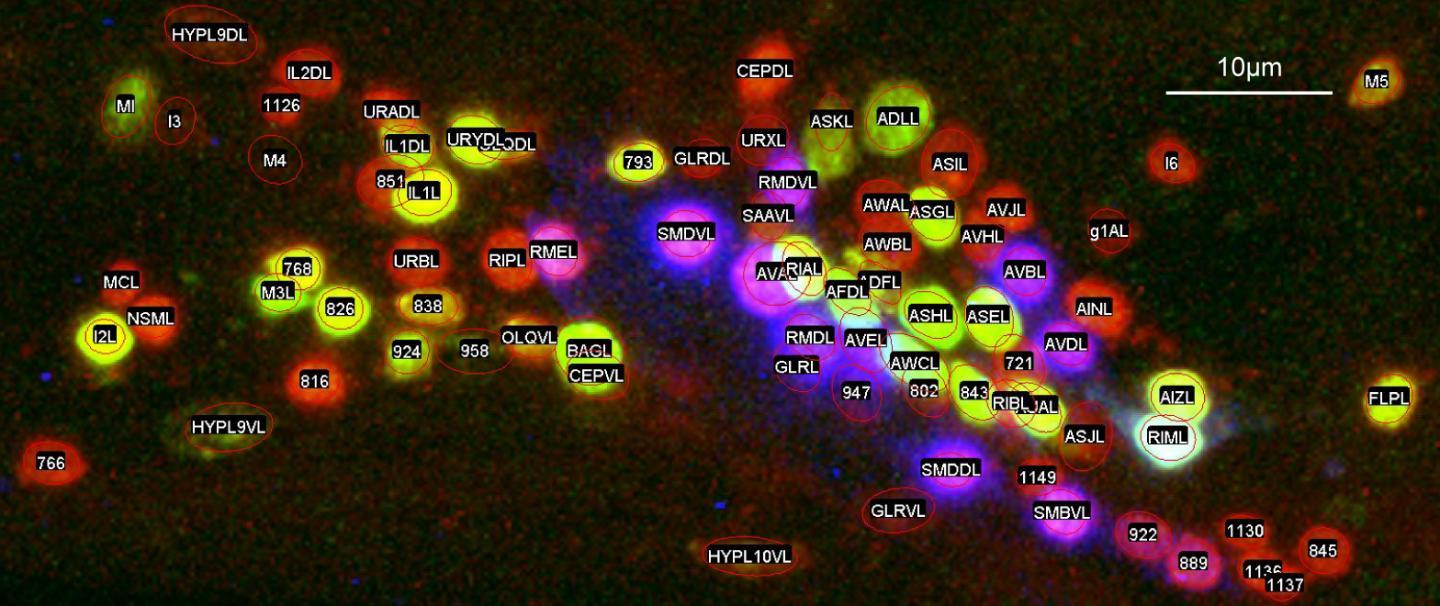 {module INSIDE STORY}
{module INSIDE STORY}
Despite their value, generalized brain atlases, so-called connectome maps, are still no help for identifying neurons in individual, live, wriggling worms.
"Imagine if you knew the names of all the cities on a map, but the cities moved each time you looked. That is what it's like, trying to compare current brain atlases to living organisms," said Professor Yuichi Iino from the University of Tokyo, co-last author of the recent research paper published in BMC Biology.
Iino's research group wants to identify and map each nerve cell in living C. elegans so that they can chart the pathways of electrical impulses that make behaviors, learning, and memory possible.
C. elegans brain neurons are not trapped in a skull, but just form a loosely packed group of 150 neurons in the head region of the animal.
"The neurons are tiny, and in the head of C. elegans they are surrounding this large bulb that's part of the digestive system, so they get pushed and pulled around a lot like the animal moves or eats," explained Iino.
Researchers began by finding unique combinations of genes that, when artificially attached to fluorescent protein tags, would cause 35 different small groups of neurons to glow under a microscope.
These new genetically modified strains of C. elegans made all of the researchers' subsequent image studies and supercomputer programming work possible.
Researchers identified individual neurons in 311 worms in total, about 10 worms for each of the 35 neuron groups, and measured the distances and relative positions between pairs of neurons in the microscopy images.
Although neurons were known to shift within each worm, no one expected the neurons to have different "home base" locations in different individuals. The positions of the central cell body of some neurons can vary by more than 0.02 millimeters between different animals, a significant distance for an animal only 1-millimeter long.
"Individual C. elegans are thought to be uniform because they all have almost the same cell lineages and a stereotyped neural circuit. It was really surprising, though, how large the positional differences are between individual animals," said Assistant Professor Yu Toyoshima, a co-first author of the recent research paper and member of the Iino lab.
The research team then used their new position variation data and the C. elegans connectome brain atlas to develop a computer program to automatically identify neurons. The program uses a mathematical algorithm to analyze a microscopy image of the C. elegans brain and assign the statistically most likely identity to each neuron based on that neuron's position in relation to other neurons.
"The algorithm is only 60 percent accurate, which is too low for fully automatic cell identification, but it speeds up our work enough to make other projects possible to understand neural networks based on whole-brain imaging data," said Toyoshima.
Part of what made this project possible in C. elegans is that every neuron was already known and named. Using a similar technique in other animals would require fine-tuned genetic manipulation to cause groups of neurons to glow under a microscope and knowing how many neurons need to be identified. 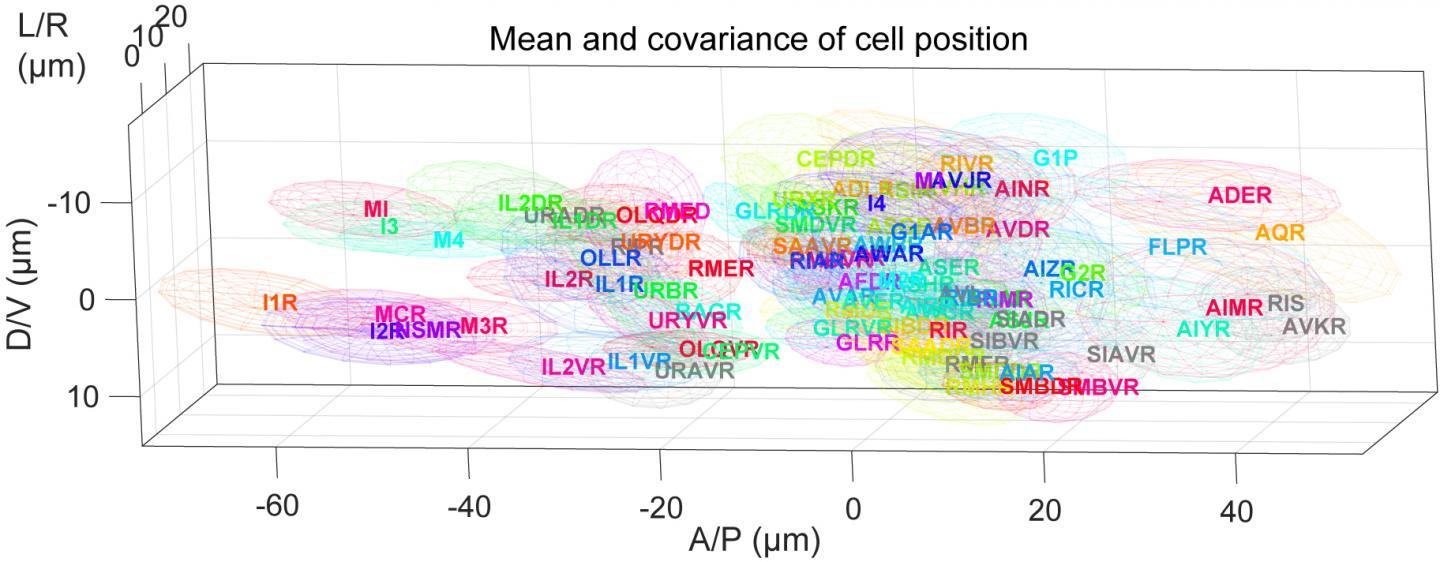
{module INSIDE STORY}
"The human brain has billions of neurons, so understanding our own brains at the single-cell level would be extremely difficult. C. elegans have small brains, but they can still learn and change behaviors, so they could allow us to understand how networks of neurons create behavior," said Iino.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?