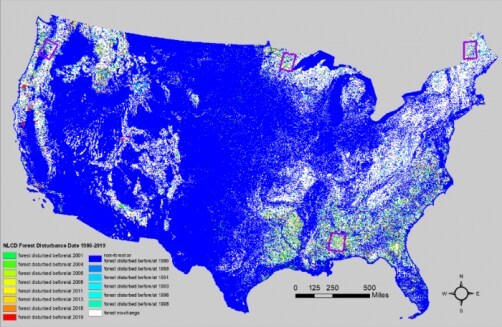
Every second, the planet loses a stretch of forest equivalent to a football field due to logging, fires, insect infestation, disease, wind, drought, and other factors. In a recently published study, researchers from the U.S. Geological Survey Earth Resources Observation and Science (EROS) Center presented a comprehensive strategy to detect when and where forest disturbance happens at a large scale and provide a deeper understanding of forest change.
“Our strategy leads to more accurate land cover mapping and updating,” said Suming Jin, a physical scientist with the EROS Center.
To understand the big picture of a changing landscape, scientists rely on the National Land Cover Database, which turns Earth-observation satellite (Landsat) images into pixel-by-pixel maps of specific features. Between 2001 and 2016, the database showed that nearly half of the land cover change in the contiguous United States involved forested areas.
“To ensure the quality of National Land Cover Database land cover and land cover change products, it is important to accurately detect the location and time of forest disturbance,” said Jin.
Jin and their team developed a method to detect forest disturbance by year. The approach combines strengths from a time-series algorithm and a 2-date detection method to improve large-region operational mapping efficiency, flexibility, and accuracy. The new technique facilitates more effective forest management and policy, among other applications.
Landsat data have been widely used to detect forest disturbance because of their long history, high spatial and radiometric resolutions, free and open data policy, and suitability for creating continental or even global mosaic images for different seasons.
“We need algorithms that can create consistent large-region forest disturbance maps to assist in producing a multi-epoch National Land Cover Database,” said Jin. “We also need those algorithms to be scalable so we can track forest change over longer periods of time.”
A commonly employed method called “2-date forest change detection” involves comparing images from two different dates while the “time-series algorithm” can provide observations for yearly or even monthly Landsat time series.
In general, 2-date forest change detection algorithms are more flexible than time-series methods and use richer spectral information. The 2-date method can easily determine changes between image bands, indices, classifications, and combinations and, therefore, detect forest disturbances more accurately. However, the 2-date method only detects changes for one time period and usually requires additional information or further processing to separate forest changes from other land cover changes.
On the other hand, time-series-based forest change detection algorithms can use spectral and long-term temporal information and produce changes for multiple dates simultaneously. However, these methods usually require every step of the time series algorithm to be processed again when a new date is added, which can be cumbersome for continuous monitoring updates and lead to inconsistencies.
Previous studies proposed ensemble approaches to improve forest change mapping accuracy, including “stacking,” or combining the output of different mapping methods. While stacking reduces omission and commission error rates, the method is computationally intensive and requires reference data for training.
Jin and team’s approach combined strengths from 2-date change detection methods and the continuous time-series change detection method, which was called the Time-Series method Using Normalized Spectral Distance (NSD) index (TSUN), to improve large-region operational mapping efficiency, flexibility, and accuracy. Using this combination, the researchers produced the NLCD 1986–2019 forest disturbance product, which shows the most recent forest disturbance date between the years 1986 and 2019 for every two-to-three-year interval.
“The TSUN index detects multi-date forest land cover changes and was shown to be easily extended to a new date even when new images were processed in a different way than previous date images,” Jin said.
The research team plans to improve the tool by increasing the time frequency and producing an annual forest disturbance product from 1986 to the present.
“Our ultimate goal is to automatically produce forest disturbance maps with high accuracy with the capability of continually monitoring forest disturbance, hopefully in real-time,” Jin said.
This work was supported by the USGS-NASA Landsat Science Team Program for Toward Near Real-time Monitoring and Characterization of Land Surface Change for the Conterminous US.
Other contributors include Jon Dewitz from the U.S. Geological Survey Earth Resources Observation and Science (EROS) Center; Congcong Li with ASRC Federal Data Solutions; Daniel Sorenson with the U.S. Geological Survey; Zhe Zhu with the University of Connecticut; Md Rakibul Islam Shogib, Patrick Danielson and Brian Granneman from KBR; Catherine Costello with the U.S. Geological Survey, Geosciences, and Environmental Change Science Center; Adam Case with Innovate! Inc.; and Leila Gass with the U.S. Geological Survey, Western Geographic Science Center.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?