A new form of magnetic interaction which pushes a formerly two-dimensional phenomenon into the third dimension could open up a host of exciting new possibilities for data storage and advanced supercomputing, scientists say.
In a new paper published today in the journal Nature Materials, a team led by physicists from the University of Glasgow describe how they have been found a new way to successfully pass information from a series of tiny magnets arrayed on an ultrathin film across to magnets on a second film below.
Their breakthrough adds both a literal and metaphorical extra dimension to ‘spintronics’, the field of science dedicated to data storage, retrieval, and processing, which has already had a major impact on the tech industry.
Anyone who’s ever played with a pair of magnets understands that opposites attract – the south pole of one magnet attracts the north pole of the other. While that’s true at the scale most people are familiar with, the way magnets interact with each other undergoes some significant changes as magnets shrink. {module In-article}
At the nanoscale – where magnetic materials can be just a few billionths of a meter in size - magnets interact with each other in strange new ways, including the possibility of attracting and repelling each other at 90-degree angles instead of straight-on.
Scientists have already learned how to exploit those unusual properties to encode and process information in thin films covered in a single layer of nanoscale magnets.
The benefits of these ‘spintronic’ systems – low power consumption, high storage capacity, and greater robustness - have made invaluable additions to technology such as magnetic hard disk drives, and won the discoverers of spintronics a Nobel prize in 2007.
However, the functionality of magnetic systems used today in computers remains confined to one plane, limiting their capacity. Now, the University of Glasgow-led team – along with partners from the Universities of Cambridge and Hamburg, the Technical University of Eindhoven and the Aalto University School of Science – have developed a new way to communicate information from one layer to another, adding the new potential for storage and computation.
Dr Amalio Fernandez-Pacheco, an EPSRC Early Career Fellow in the University’s School of Physics and Astronomy, is the lead author on the paper. He said: “The discovery of this new type of interaction between neighbor layers gives us a rich and exciting way to explore and exploit unprecedented 3D magnetic states in multi-layered nanoscale magnets.
“It’s a bit like being given an extra note in a musical scale to play with - it opens up a whole new world of possibilities, not just for conventional information processing and storage, but potentially for new forms of computing we haven’t even thought of yet.”
The inter-layer transmission of information the team has created relies on what is known to physicists as chiral spin interactions, a type of magnetic force that favors a particular sense of rotation in neighbor nanoscale magnets. Thanks to recent advances in spintronics, it is now possible to stabilize these interactions within a magnetic layer. This has for instance been exploited to create skyrmions, a type of nanoscale magnetic object with superior properties for computing applications.
The team’s research has now extended these types of interactions to neighboring layers for the first time. They fabricated a multi-layered system formed by ultra-thin magnetic films separated by non-magnetic metallic spacers. The structure of the system and precise tuning of the properties of each layer and its interfaces creates unusual canted magnetic configurations, where the magnetic field of the two layers forms angles between zero and 90 degrees.
{module In-article}
Unlike in standard multi-layered magnets, it becomes easier for these magnetic fields to form clockwise configurations than anticlockwise ones, a fingerprint that an interlayer chiral spin interaction exists in between the two magnetic layers. This breaking of rotational symmetry was observed at room temperature and under standard environmental conditions. As a result, this new type of interlayer magnetic interaction opens exciting perspectives to realize topologically complex magnetic 3D configurations in spintronic technologies.
The team’s paper, titled ‘Symmetry-Breaking Interlayer Dzyaloshinskii-Moriya Interactions in Synthetic Antiferromagnets’, is published in Nature Materials. The research was funded by the Engineering and Physical Sciences Research Council.
A tweaked gene or two among the millions or even billions of proteins that make up an organism's DNA are often all that distinguish the drought-tolerant plant or the person pre-disposed to cancer.
That's why a better understanding of genetic variation within a species could, among other things, help improve the selection of crops for local conditions and detection of disease, according to Joann Mudge, a senior research scientist at the nonprofit National Center for Genome Resources.
A generation ago, recording an organism's DNA from beginning to end was so laborious and expensive that scientists celebrated when they completed the task for a single bacterium. But as genome sequencing becomes faster and cheaper, scientists increasingly have access to insights about which genes do what, Mudge said.
"We're sequencing multiple individuals of some species," including plants and other complex organisms, Mudge said. That allows scientists to begin to sort out which segments of DNA from a species' core genome and which correspond to traits shared by only some individuals, she said.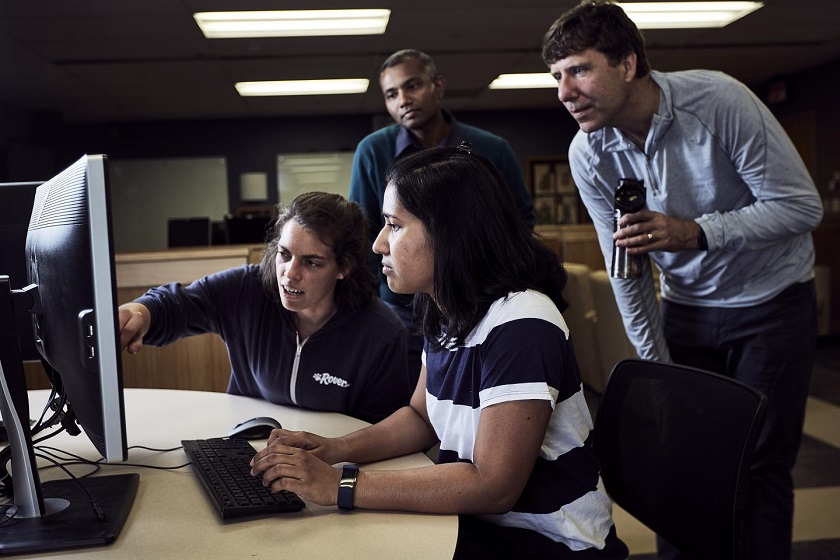  {module In-article}
{module In-article}
But the growing field of pangenomics, as it is called, presents a major analytical challenge. That's why NCGR recently partnered with Montana State University computer scientists to develop software that can compare multiple genomes and make sense of the results. The project is backed by a three-year, $662,000 grant from the National Science Foundation.
"We've been very happy with the way it's working," said Brendan Mumey, a professor in the Gianforte School of Computing in MSU's Norm Asbjornson College of Engineering. He and Mudge are co-leading the project.
According to Mumey, previously available software struggled with analyzing pangenomes for relatively primitive organisms such as the common yeast Saccharomyces cerevisiae, whose genome contains only 12 million of the DNA units known as base pairs. (By comparison, the human genome contains 3 billion base pairs.) Among the known strains of the yeast, minor genetic variations account for physical adaptations such as the ability of brewer's yeast to survive alcohol during the making of beer and wine.
"It's a classic 'big data' problem," Mumey said, referring to the field of supercomputing that deals with exceptionally large and complex data sets.
MSU assistant professor of computer science Indika Kahanda, a member of the research team, specializes in developing the "machine learning" models that help the new software adjust its gene-sorting analysis according to input from scientists. That approach has helped the team, which includes NCGR research scientist Thiru Ramaraj, identify genes of interest in a yeast pangenome that includes roughly 100 strains. Ramaraj earned his doctorate in computer science in 2010 at MSU, where Mumey was his adviser.
Mumey said the researchers' next step is to continue to refine the software so it can handle larger and more complex genomes, such as those of plants. The computational techniques being used "are still in their infancy," he said.
Eventually, pangenomics could help medical professionals diagnose a variety of diseases that have a genetic component, Mudge said. Most inherited breast cancer can be traced to mutations in just two genes, but other genetic diseases are thought to stem from more complex changes across larger areas of DNA.
The improved pangenomics tool is already helping scientists break out of a mold of comparing genomes to a single, arbitrary reference, Mudge said. Instead, researchers can represent a species' entire genome with all its nuance and variety.
"It's a hard problem to solve," Mudge said. "This has been a great collaboration."


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module In-article}
{module In-article} {module In-article}
{module In-article}