A new approach brings the hope of new therapeutic options for suppressing seasonal influenza and avian flu: On the basis of an empty – and therefore non-infectious – the shell of a phage virus, researchers from Berlin have developed a chemically modified phage capsid that “stifles” influenza viruses. Perfectly fitting binding sites cause influenza viruses to be enveloped by the phage capsids in such a way that it is practically impossible for them to infect lung cells any longer. This phenomenon has been proven in pre-clinical trials, also involving human lung tissue. Researchers from the Leibniz-Forschungsinstitut für Molekulare Pharmakologie (FMP), Freie Universität Berlin (FU), Technische Universität Berlin (TU), Humboldt-Universität (HU), the Robert Koch Institute (RKI) and Charité were involved in this groundbreaking work. The results are also being used for the immediate investigation of the coronavirus. The findings have now been published in an academic journal.
Influenza viruses are still highly dangerous: The World Health Organization (WHO) estimates that flu is responsible for up to 650,000 deaths per year worldwide. Current antiviral drugs are only partially effective because they attack the influenza virus after lung cells have been infected. It would be desirable – and much more effective – to prevent infection in the first place.
This is exactly what the new approach from Berlin promises. The phage capsid, developed by a multidisciplinary team of researchers, envelops flu viruses so perfectly that they can no longer infect cells. “Pre-clinical trials show that we are able to render harmless both seasonal influenza viruses and avian flu viruses harmless with our chemically modified phage shell,” explained Professor Dr. Christian Hackenberger, Head of the Department Chemical Biology at the Leibniz-Forschungsinstitut für Molekulare Pharmakologie (FMP) and Leibniz-Humboldt Professor for Chemical Biology at HU Berlin. “It is a major success that offers entirely new perspectives for the development of innovative antiviral drugs.” 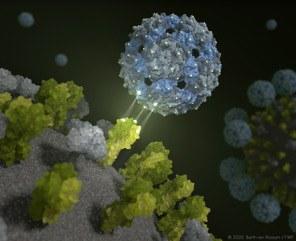 {module INSIDE STORY}
{module INSIDE STORY}
Multiple bonds fit like hook-and-loop tape
The new inhibitor makes use of a feature that all influenza viruses have: There are trivalent receptors on the surface of the virus, referred to as hemagglutinin protein, that attaches to sugar molecules (sialic acids) on the cell surface of lung tissue. In the case of infection, viruses hook into their victim – in this case, lung cells – like a hook-and-loop fastener. The core principle is that these interactions occur due to multiple bonds, rather than single bonds.
It was the surface structure of flu viruses that inspired the researchers to ask the following initial question more than six years ago: Would it not be possible to develop an inhibitor that binds to trivalent receptors with a perfect fit, simulating the surface of lung tissue cells?
We now know that this is indeed possible – with the help of a harmless intestinal inhabitant: The Q-beta phage has the ideal surface properties and is excellently suited to equip it with ligands – in this case, sugar molecules – as “bait”. An empty phage shell does the job perfectly. “Our multivalent scaffold molecule is not infectious, and comprises 180 identical proteins that are spaced out exactly as the trivalent receptors of the hemagglutinin on the surface of the virus,” explained Dr. Daniel Lauster, a former Ph.D. student in the Group of Molecular Biophysics (HU) and now a postdoc at FU Berlin. “It, therefore, has the ideal starting conditions to deceive the influenza virus – or, to be more precise, to attach to it with a perfect spatial fit. In other words, we use a phage virus to disable the influenza virus!”
To enable the Q-beta scaffold to fulfill the desired function, it must first be chemically modified. Produced from E. coli bacteria at TU Berlin, Professor Hackenberger’s research group at FMP and HU Berlin use synthetic chemistry to attach sugar molecules to the defined positions of the virus shell.
The virus is deceived and enveloped
Several studies using animal models and cell cultures have proven that the suitably modified spherical structure possesses considerable bond strength and inhibiting potential. The study also enabled the Robert Koch Institute to examine the antiviral potential of phage capsids against many current influenza virus strains, and even against avian flu viruses. Its therapeutic potential has even been proven on human lung tissue, as fellow researchers from the Medical Department, Division of Infectiology and Pneumology, of Charité were able to show: When tissue infected with flu viruses was treated with the phage capsid, the influenza viruses were practically no longer able to reproduce.
The results are supported by structural proof furnished by FU scientists from the Research Center of Electron Microscopy (FZEM): High-resolution cryo-electron microscopy and cryo-electron microscopy show directly and, above all, spatially, that the inhibitor completely encapsulates the virus. In addition, mathematical-physical models were used to simulate the interaction between influenza viruses and the phage capsid on the supercomputer. “Our computer-assisted calculations show that the rationally designed inhibitor does indeed attach to the hemagglutinin, and completely envelops the influenza virus,” confirmed Dr. Susanne Liese from the AG Netz of FU Berlin. “It was therefore also possible to describe and explain the high bond strength mathematically.”
Therapeutic potential requires further research
These findings must now be followed up by more preclinical studies. It is not yet known, for example, whether the phage capsid provokes an immune response in mammals. Ideally, this response could even enhance the effect of the inhibitor. However, it could also be the case that an immune response reduces the efficacy of phage capsids in the case of repeated-dose exposure, or that flu viruses develop resistances. And, of course, it has yet to be proven that the inhibitor is also effective in humans.
Nonetheless, the alliance of Berlin researchers is certain that the approach has great potential. “Our rationally developed, three-dimensional, multivalent inhibitor points to a new direction in the development of structurally adaptable influenza virus binders. This is the first achievement of its kind in multivalency research,” emphasized Professor Hackenberger. The chemist believes that this approach, which is biodegradable, non-toxic and non-immunogenic in cell culture studies, can in principle also be applied to other viruses, and possibly also to bacteria. It is evident that the authors regard the application of their approach to the current coronavirus as one of their new challenges. The idea is to develop a drug that prevents coronaviruses from binding to host cells located in the throat and subsequent airways, thus preventing infection.
Berlin university alliance at its best
Cooperation between scientists from different disciplines played a major role in the discovery of the new influenza inhibitor. Biologists, chemists, physicists, virologists, medical scientists and imaging specialists from three Berlin universities HU, FU and TU, the Robert Koch Institute, Charité and, last but not least, FMP were all involved in the project. “In my opinion, such a complex project could only have been undertaken in Berlin, where there truly are experts for every issue,” stated Professor Dr. Andreas Herrmann, Head of Molecular Biophysics at HU Berlin. “It was the Berlin university alliance at its best,” he added, “and I hope that the follow-up studies will be equally successful.”
The project was funded within Collaborative Research Center 765 (Speaker Professor Dr. Rainer Haag, FU Berlin) of the German Research Foundation (DFG).


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module INSIDE STORY}
{module INSIDE STORY} {module INSIDE STORY}
{module INSIDE STORY}
 {module INSIDE STORY}
{module INSIDE STORY}