New supercomputer modeling could play a big part in exit strategies and lifting air travel restrictions
Millions of more people across the EU could have contracted COVID-19 had strict international travel bans not been implemented, shows a new report by supercomputer modeling experts at Stanford University.
Using a newly developed mathematical epidemiology simulation, the study, published in Computer Methods in Biomechanics and Biomedical Engineering, predicts the huge impact that limiting air travel across the 27 EU nations had on restricting the spread of the disease. 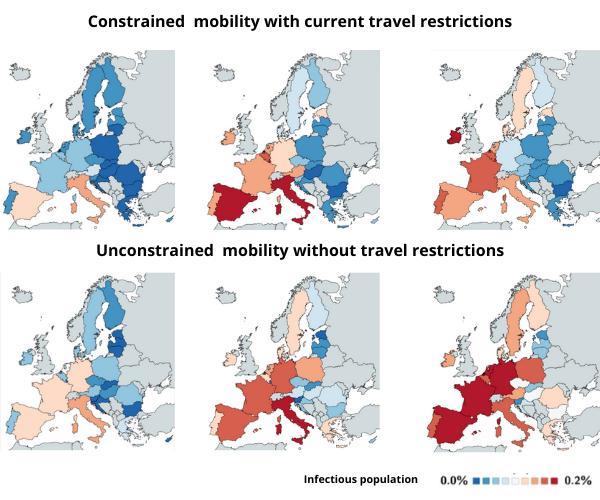 {module INSIDE STORY}
{module INSIDE STORY}
The simulation can show live estimated figures for the growth of spread for each country if we were to remove travel bans today. The images above show how 0.2% of some populations could have become infected by 20th April (when the study was written, 5 April), however, these figures change daily.
This new model could now play a vital part in establishing politicians' exit strategies, with the team able to virtually lift travel restrictions between individual communities, states, or countries, to explore the potential gradual changes in spreading patterns and outbreak dynamics.
"There is a well-reasoned fear that easing of current (travel restriction) measures, even slightly, could trigger a new outbreak and accelerate the spread to an unmanageable degree," lead author Ellen Kuhl, Professor of Mechanical Engineering at Standford University comments.
"Global network mobility models, combined with local epidemiology models, can provide valuable insight into different exit strategies. Our results demonstrate that mathematical modeling can provide guidelines for political decision making with the ultimate goal to gradually return to normal while keeping the rate of new COVID-19 infections steady and manageable," says Kevin Linka, lead author and a postdoctoral researcher in Dr. Kuhl's group.
From its European origin in Italy, the novel coronavirus spread rapidly via the strongest network connections to Germany, Spain, and France, while slowly reaching the less connected countries, Estonia, Slovakia, and Slovenia.
Currently, the levels of the population known to be infected with the disease vary from country to country, however as of April 18, with the flight being reduced by 89% in Germany, 93% in France, 94% in Italy, and 95% in Spain (Eurostat 2020), the graphs in this study show how the spread has been contained.
"Strikingly, our results suggest that the emerging pattern of the COVID-19 outbreak closely followed global mobility patterns of air passenger travel," confirms Professor Kuhl, whose model can also predict the emerging global diffusion pattern of a pandemic at the early stages of the outbreak.
"Our results suggest that unconstrained mobility would have significantly accelerated the spreading of COVID-19, especially in Central Europe, Spain, and France."
Unfortunately, the model also confirms how travel bans were introduced too late to stop the Europe-wide outbreak altogether.
"A recent study based on a global metapopulation disease transmission model for the COVID-19 outbreak in China has shown that the Wuhan travel ban essentially came too late, at a point where most Chinese cities had already received many infected travelers (Chinazzi et al. 2020). Our study shows a similar trend for Europe, where travel restrictions were only implemented a week after every country had reported cases of COVID-19 (European Centre for Disease Prevention and Control 2020).
"As a natural consequence, unfortunately, no European country was protected from the outbreak," Professor Kuhl, who is the Robert Bosch Chair of Mechanical Engineering at Standford added.
The first official case of COVID-19 in Europe was reported in France on January 24, 2020, followed by Germany and Finland only three and five days later. Within only six weeks, all 27 countries of the European Union were affected, with the last cases reported in Malta, Bulgaria, and Cyprus on March 9, 2020. At this point, there were 13,944 active cases within the European Union and the number of active cases doubled every three to four days (European Centre for Disease Prevention and Control 2020).
Dr. Kuhl adds that although air travel is certainly not the only determinant of the outbreak dynamics, their findings indicate that "mobility is a strong contributor to the global spreading of COVID-19". This is becoming especially important now that many countries are beginning to lift their travel restrictions in an attempt to gradually return to normal.
Other limitations highlighted - like any infectious disease model - include the simulation being subject to data uncertainties from differences in testing, inconsistent diagnostics, incomplete counting, and delayed reporting across all countries.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module INSIDE STORY}
{module INSIDE STORY} 