UK scientists have developed a pioneering new technique that harnesses the cutting-edge capabilities of AI to model and map the natural environment in intricate detail.
A team of experts, including Charlie Kirkwood from the University of Exeter, has created a sophisticated new approach to modeling the Earth’s natural features in greater detail and accuracy.
The new technique can recognize intricate features and aspects of the terrain far beyond the capabilities of more traditional methods and use these to generate enhanced-quality environmental maps.
Crucially, the new system could also pave the way to unlocking discoveries of the relationships within the natural environment, that may help tackle some of the greater climate and environment issues of the 21st century.
The study is published in the academic journal Mathematical Geosciences, as part of a special issue on geostatistics and machine learning.
Modeling and mapping the environment is a lengthy, time-consuming, and expensive process. Cost limits the number of observations that can be obtained, which means that creating comprehensive spatially-continuous maps depends upon filling in the gaps between these observations.
Scientists can use a range of information sources to help fill in these observation gaps, such as terrain elevation data and satellite imagery. However, conventional modeling methods rely on users to manually engineer predictive features from these datasets – for example generating slope angles and curvatures from terrain elevation data in the hope that these can help explain the spatial distribution of the variable being mapped.
However, scientists believe there are likely to be many more nuanced relationships at play within the natural environment that models based on traditional manual feature-engineering approaches may simply miss.
The pioneering new AI approach, developed in the study, poses environmental information extraction as an optimization problem. Doing so allows it to automatically recognize and make use of relationships that may otherwise go unnoticed and unutilized by humans using more traditional modeling methods.
In addition to improving map quality, this also unlocks the potential for the discovery of new relationships in the natural environment by AI, while simultaneously eliminating huge amounts of trial-and-error experimentation in the modeling process.
Charlie Kirkwood, a postgraduate student at the University of Exeter said: “To be useful for decision making, we need our models to provide answers that are as specific as possible while also being trustworthy – and that means creating accurate measures of the uncertainty associated with our estimates, which in this case are predictions at unmeasured locations.”
“Our AI approach is set within a Bayesian statistical framework which allows us to quantify these uncertainties and provide a range of uncertainty measures, including credible intervals, exceedance probabilities, and other more bespoke products that will feed directly into decision-making processes. Crucially, all this is provided whilst harnessing any available information more effectively than traditional approaches allow – which you can see coming through in the detail of the map“
The new approach was demonstrated using stream sediment calcium concentration observations from the British Geological Survey’s Geochemical Baseline Survey of the Environment (G-BASE) project.
The distribution of calcium in the environment, which has standalone importance for its impact on soil fertility, is controlled primarily by geology – with different rock types containing different proportions of calcium – but also by hydrological processes at the surface.
Calcium, therefore, provides a challenging use case for the AI approach, which must learn to recognize and utilize features relating to both bedrock geology (e.g. differing terrain textures, breaks of slope) and surface hydrology (e.g. drainage, river channels).
The method, the scientists say, has produced a spectacularly detailed and accurate map which, despite depicting just one element – calcium, reveals the geology of Britain in arguably a new level of detail thanks to the information-extracting power of the new AI approach. The team believes that by combining the research skills, expertise, and data resources of its partners - the University of Exeter, Met Office, and British Geological Survey - this work presents a new dawn for environmental mapping practices in the age of AI.
Professor Gavin Shaddick, from the University of Exeter, added “This is a fantastic example of Environmental Intelligence, the use of AI to help solve challenges in environmental science. This work is an exemplar in integrating technical knowledge of AI and machine learning with expertise in geosciences to produce a new methodology that directly addresses crucial questions in mapping environmental information. The resulting methodological advances could be used to produce detailed maps of a wide variety of environmental hazards and have the potential to provide a rich source of information for both scientists and decision-makers.”
Garry Baker, Interim Chief Digital Officer, British Geological Survey added: “This paper is an excellent demonstration of how environmental information such as the BGS geochemical database can be re-assessed via new approaches (AI spatial interpolation). It exemplifies the benefits of ongoing environmental research and how this can draw upon the extensive datasets available to everyone through the National Geoscience Data Centre and wider NERC, and UKRI data repositories.”
Dr. Kirstine Dale, the Met Office’s Principal Fellow for Data Science and Co-Director for Joint Centre for Excellence in Environmental Intelligence commented on the value of this work: “This is an important example of how data science has the potential to transform our understanding of the natural world. Critically, it highlights what can be achieved by working across disciplines, in this case bringing together mathematicians, weather specialists and computer scientists enrich our knowledge of the natural world in a way that no single discipline can.”


 How to resolve AdBlock issue?
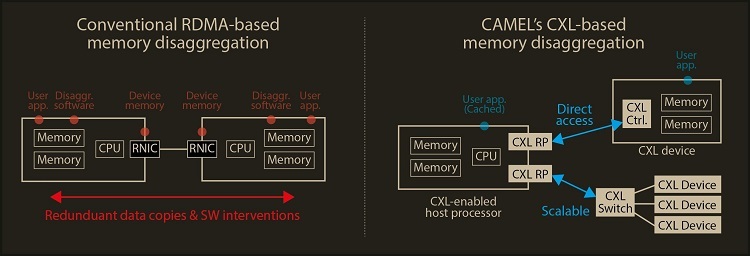
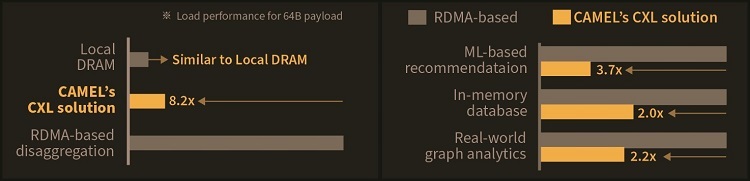
How to resolve AdBlock issue?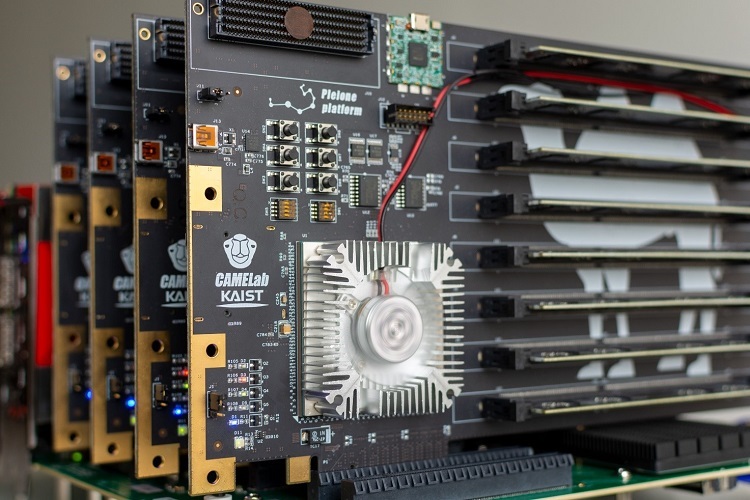  A KAIST team compute express link (CXL) provides new insights on memory disaggregation and ensures direct access and high-performance capabilities
A KAIST team compute express link (CXL) provides new insights on memory disaggregation and ensures direct access and high-performance capabilities