Birdsong has long captivated bird enthusiasts and ecologists, providing valuable insight into avian behavior, preferred habitats, and species identification. However, identifying rare bird species through their songs has been a significant challenge for researchers and conservationists. To address this issue, a team of researchers at the University of Moncton in Canada has developed a groundbreaking deep-learning AI tool called ECOGEN. This innovative tool creates lifelike birdsongs, enhancing samples of underrepresented species. It's revolutionizing efforts in bird monitoring and conservation.
Improving Birdsong Identification with AI
In recent years, several phone apps and software have emerged, enabling ecologists and the public to identify common bird species through their songs. These tools have proved useful for widely recognized species. However, they often face challenges with rare or elusive birds, of which there are limited recorded samples. ECOGEN aims to bridge this gap by generating artificial birdsong samples that can be used to train audio identification tools for ecological monitoring.
Dr. Nicolas Lecomte, one of the lead researchers, explains the significance of ECOGEN: "Due to significant global changes in animal populations, there is an urgent need for automated tools such as acoustic monitoring to track shifts in biodiversity. However, the AI models used to identify species in acoustic monitoring lack comprehensive reference libraries. ECOGEN addresses this gap by creating new instances of bird sounds to support AI models. This expands the sound library for species with limited recordings, ultimately benefiting conservation efforts."
The Power of Synthetic Bird Songs
Researchers at the University of Moncton have demonstrated that incorporating artificial birdsong samples generated by ECOGEN significantly improves bird song classification accuracy. By adding these lifelike sounds to the existing audio identification tools, researchers observed an average increase of 12% in classification accuracy. This breakthrough has tremendous implications for ecological monitoring, enabling researchers to more effectively track shifts in biodiversity and identify rare and endangered bird species.
Moreover, ECOGEN has the potential to contribute to the conservation of endangered bird species by providing valuable insights into their vocalizations, behaviors, and habitat preferences. The tool offers a unique opportunity to study and understand rare birds that are difficult to observe in the wild, furthering our efforts to protect and preserve these vulnerable species.
ECOGEN is an AI-powered tool that was initially developed for bird species but has the potential to be adapted for other animals such as mammals, fish, insects, and amphibians. Dr. Lecomte, the creator of ECOGEN, believes that this versatility opens up new avenues for ecological research and monitoring, enabling scientists to gain a deeper understanding of diverse animal populations and their acoustic behaviors.
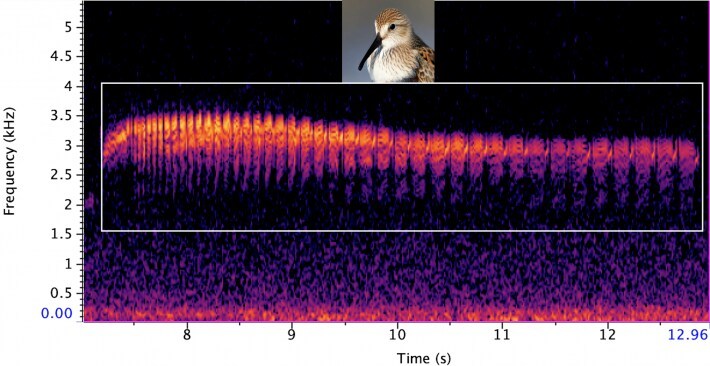
The tool employs a sophisticated process to generate lifelike spectrograms, which are visual representations of sounds. It converts real recordings of bird songs into spectrograms, generates new AI images to expand the dataset for rare species with limited recordings, and converts the AI-generated images into audio to train bird sound identifiers. To develop ECOGEN, the researchers utilized a dataset of 23,784 wild bird recordings from around the world, encompassing 264 species. This vast dataset allowed the AI tool to learn and generate accurate representations of various bird species, enhancing the effectiveness of audio identification tools used in ecological monitoring.
ECOGEN is open-source and can be accessed and used by researchers and conservationists worldwide. It is designed to function on basic computers, making it accessible to those with limited resources. This breakthrough technology has the potential to revolutionize bird monitoring efforts, aiding in the conservation of endangered species and providing valuable insights into their vocalizations and behaviors.
As further research is conducted, ECOGEN has the potential to expand its applications to other animal groups, including mammals, fish, insects, and amphibians. By providing a comprehensive and accessible AI tool for acoustic monitoring, scientists can gain a deeper understanding of our natural world and contribute to the preservation of biodiversity.
In conclusion, ECOGEN represents a significant advancement in the field of ecological monitoring and conservation. By generating lifelike birdsongs, this powerful AI tool enhances the accuracy of bird song identification and expands our understanding of rare and endangered species. With its open-source nature and potential for further adaptation, ECOGEN offers a promising future for conservation efforts worldwide.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?