Underwater robots are being widely used as tools in a variety of marine tasks. The RobDact is one such bionic underwater vehicle, inspired by a fish called Dactylopteridae known for its enlarged pectoral fins. A research team has combined computational fluid dynamics and a force measurement experiment to study the RobDact, creating an accurate hydrodynamic model of the RobDact that allows them to better control the vehicle. 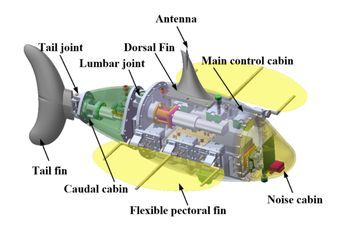
The team published their findings in Cyborg and Bionic Systems on May 31, 2022.
Underwater robots are now used for many marine tasks, including in the fishery industry, underwater exploration, and mapping. Most traditional underwater robots are driven by a propellor, which is effective for cruising in open waters at a stable speed. However, underwater robots often need to be able to move or hover at low speeds in turbulent waters, while performing a specific task. It is difficult for the propellor to move the robot in these conditions. Another factor when an underwater robot is moving at low speeds in unstable flowing waters is the propeller’s “twitching” movement. This twitching generates unpredictable fluid pulses that reduce the robot’s efficiency.
In recent years, researchers have worked to create underwater robots that mimic living creatures. These bionic vehicles move through the water similar to the ways fish or manta rays move. Compared with traditional underwater propulsion vehicles, these bionic underwater vehicles operate more efficiently and robustly in the water, while being environmentally friendly.
Underwater robots are affected by the surrounding fluid as they move through the water. This phenomenon is called the hydrodynamic effect. While moving in the water, the robot must deal with unknown water flow and force, which can cause unnecessary changes in the robot’s position.
To better control the robot, researchers need a more accurate hydrodynamic model. Creating this model is usually very complex and difficult. In addition, the real underwater environment is changeable and difficult to predict, so the model parameters can shift with a change in the environment. Researchers have been using computational fluid dynamics to create hydrodynamic models for underwater robots. However, the models created with computational fluid dynamics alone are not as precise and practical as they need to be. To overcome this challenge, the research team tried a different approach. “To make the hydrodynamic model more accurate and practical, we combined the computational fluid dynamics and a force measurement experiment,” said Rui Wang, a researcher at the Institute of Automation, Chinese Academy of Sciences.
Using computational fluid dynamics, the researchers identified the parameters in the hydrodynamic model. Then they developed a force measurement platform to obtain the force generated by the RobDact vehicle. With this process, they could obtain both the disturbing force and the force generated by the RobDact in any complex environment. “This could help us have a better understanding of the underwater vehicle’s motion state, and control the underwater vehicle more accurately,” said Qiyuan Cao, a researcher at the Institute of Automation, Chinese Academy of Sciences.
With their experiment, the team was able to determine the hydrodynamic force of the RobDact at different speeds. The force measurement platform they developed allowed them to measure the force of RobDact in the X, Y, and Z directions. They established a mapping relationship between the RobDact fluctuation parameters and the thrust of the vehicle through their force measurement experiments. By merging the rigid body dynamic model of RobDact with the thrust mapping model, the researchers were able to develop an accurate and practical hydrodynamic model of the RobDact in varying motions.
Looking to the future, the researchers intend to study the intelligent control of bionic underwater vehicles using the hydrodynamic model in conjunction with artificial intelligence methods, such as reinforcement learning. “The ultimate goal is to promote the practical application of bionic underwater vehicles in water environment monitoring and underwater search and rescue,” said Wang.
The research team includes Qiyuan Cao and Tiandong Zhang from the Chinese Academy of Sciences, Beijing, and the University of Chinese Academy of Sciences; Rui Wang and Yu Wang from the Chinese Academy of Sciences, Beijing; and Shuo Wang from the Chinese Academy of Sciences, Beijing; the University of Chinese Academy of Sciences, and the Chinese Academy of Sciences, Shanghai.
The research is funded by the Beijing Natural Science Foundation, Beijing Nova Program, National Natural Science Foundation of China, Youth Innovation Promotion Association (Chinese Academy of Sciences), and the Young Elite Scientist Sponsorship Program (China, Association for Science and Technology).


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 
