Implications of this work include a better understanding of opportunities and appropriate tools for short-term prediction of future trends when variability is high, as well as replacement strategies for datasets with missing values.
A UCF-developed forecast model, which uses machine learning, was able to predict the spread of COVID-19 cases in Florida. Although more research is needed, the model could become useful in predicting the spread of the next big virus, which could help healthcare agencies prepare their response. 
Md Mamunur Rashid, a graduate researcher at UCF’s Virtual Readability Lab, has presented the study results alongside fellow researcher and UCF Assistant Professor Ben Sawyer ’14MS ’15PhD at the seventh North American International Conference on Industrial Engineering and Operations Management (IEOM) in Orlando June 12-14.
The meeting is a top conference for its field and is a forum for researchers, academics, and practitioners to exchange ideas and discuss recent developments within the field.
Their work predicts different parameters key to a proper COVID-19 response, through machine learning methods. If activated, a proper health response considers the planning and management of healthcare systems, determines where healthcare providers are most needed, initiates programs ensuring wellbeing, and facilitates quality education. These responses can be planned and designed only when the future can be forecasted, which can be accomplished through the models developed in this research.
The number of patients affected, hospitalizations, and deaths in the context of the state of Florida is considered to provide a blueprint for regional pandemic response.
Implications of this work include a better understanding of opportunities and appropriate tools for short-term prediction of future trends when variability is high, as well as replacement strategies for datasets with missing values. Partial, unrecorded, or non-updated data from the Florida Department of Health contributed to the missing values in the datasets. A strategy was applied to complete the partially recorded or missing data, but the data retained high variability.
“Robust predictions of COVID-19 for the future rely upon modeling of COVID data for unique regional populations,” Sawyer says. “As we transition to living alongside this virus, the present study provides important insight, characterizing how this virus interacts with the exceptional diversity of age and culture found in Florida."
How the Model Works
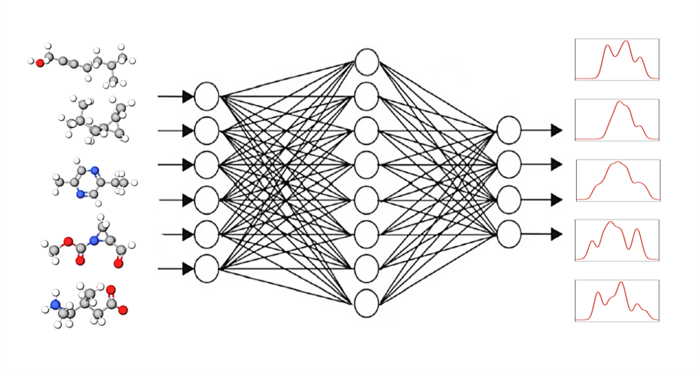
To predict these variables, 20 inputs in four categories (type of tests performed, gender, race, and age group) were collected. Official data from the Florida State Department of Health were collected and submitted to a linear regression model, fuzzy logic model, and long short-term memory (LSTM) deep learning model with the intent of producing predictions as close as possible to the actual numbers.
The mean absolute percentage error (MAPE) was calculated to measure the deviation of the predicted result from the actual values. In addition, a one-way analysis of the variance model was developed for each output parameter to statistically assess the results of these models.
The LSTM deep learning model outperformed the fuzzy model, which in turn outperformed linear regression, in terms of the MAPE for the “number of Florida residents affected.”
For the “number of patients hospitalized,” the LSTM deep learning model again outperformed the regression model, while the regression model and the fuzzy model were not significantly different.
However, no model significantly outperformed the other for “the number of patient deaths.”
“These findings suggest that in regional data, the fuzzy model outperforms linear regression, and the LSTM deep learning model outperforms both the fuzzy model and the linear regression model,” Rashid says.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?