Proteins are key molecules in living cells. They are responsible for nearly every task of cellular life and are essential for the maintenance of the structure, function, and regulation of tissues and organs in the human body. The cells in the human body can form thousands of different types of proteins (the so-called proteome), which perform a plethora of diverse functions, all crucial for cell viability and human health. Assigning functions to the vast array of proteins present in our cells remains a challenging task in cell biology.
Scientists have now produced a co-regulation map of the human proteome, which was able to capture relationships between proteins that do not physically interact or co-localize. This will enable the prediction and assignment of functions to uncharacterized human proteins. The co-regulation map can be explored at http://www.proteomeHD.net.  {module In article}
{module In article}
Many of the functions of human proteins are still unknown, but researchers at the Wellcome Centre for Cell Biology, the University of Edinburgh and the Institute of Biotechnology, Technische Universität Berlin have applied large scale quantitative proteomics and machine-learning to produce a protein covariation dataset of the human proteome. The dataset forms the basis of a co-regulation map which can be used to predict the potential function of uncharacterized human proteins.
"In this study, we took thousands of mass spectrometry experiments that other laboratories had published over the past few years and re-purposed them in a way that was completely unrelated to what the original authors had intended. We used a machine-learning algorithm to mine this huge collection of data and managed to assign a biological function to hundreds of proteins (genes) that were previously uncharacterized", said Professor Juri Rappsilber and Dr. Georg Kustatscher, of the University of Edinburgh and TU Berlin.
Exploring the map revealed unexpected co-regulation partners, namely the peroxisomal membrane protein PEX11β with mitochondrial respiration factors. In cooperation with Professor Michael Schrader and his team at the University of Exeter, this led to the identification of a novel interaction between two crucial cellular organelles - mitochondria and peroxisomes.
"Peroxisomes and mitochondria in mammals are intimately linked and cooperate in the breakdown of fatty acids and cellular energy balance. Using live-cell imaging we revealed that PEX11β mediates the formation of membrane protrusions, which emanate from peroxisomes and interact with mitochondria. They likely function in the metabolic cooperation and crosstalk between both organelles, and may facilitate the transfer of metabolites during mitochondrial energy (ATP) production", said Professor Michael Schrader, of the University of Exeter.
The scientists at the University of Edinburgh created the website http://www.proteomeHD.net to allow users to search for a protein of interest, showing its position in the co-regulation map together with any co-regulation partners.
The online map is interactive and zoomable, making it easy to explore the neighborhood of a query protein. It is designed to support researchers in exploring co-regulation data at multiple scales, to validate existing hypotheses or to create new ones.
"With an ever-increasing amount of protein expression data being made available, protein co-expression analysis has huge potential for gene function annotation. In a time when "big data" becomes more and more relevant for life science, a key lessons we learned from this project is: never throw away your data - they can be re-purposed, recycled, and with the right tools there is plenty more information and knowledge that can be extracted from them", said Dr. Georg Kustatscher, of the University of Edinburgh.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 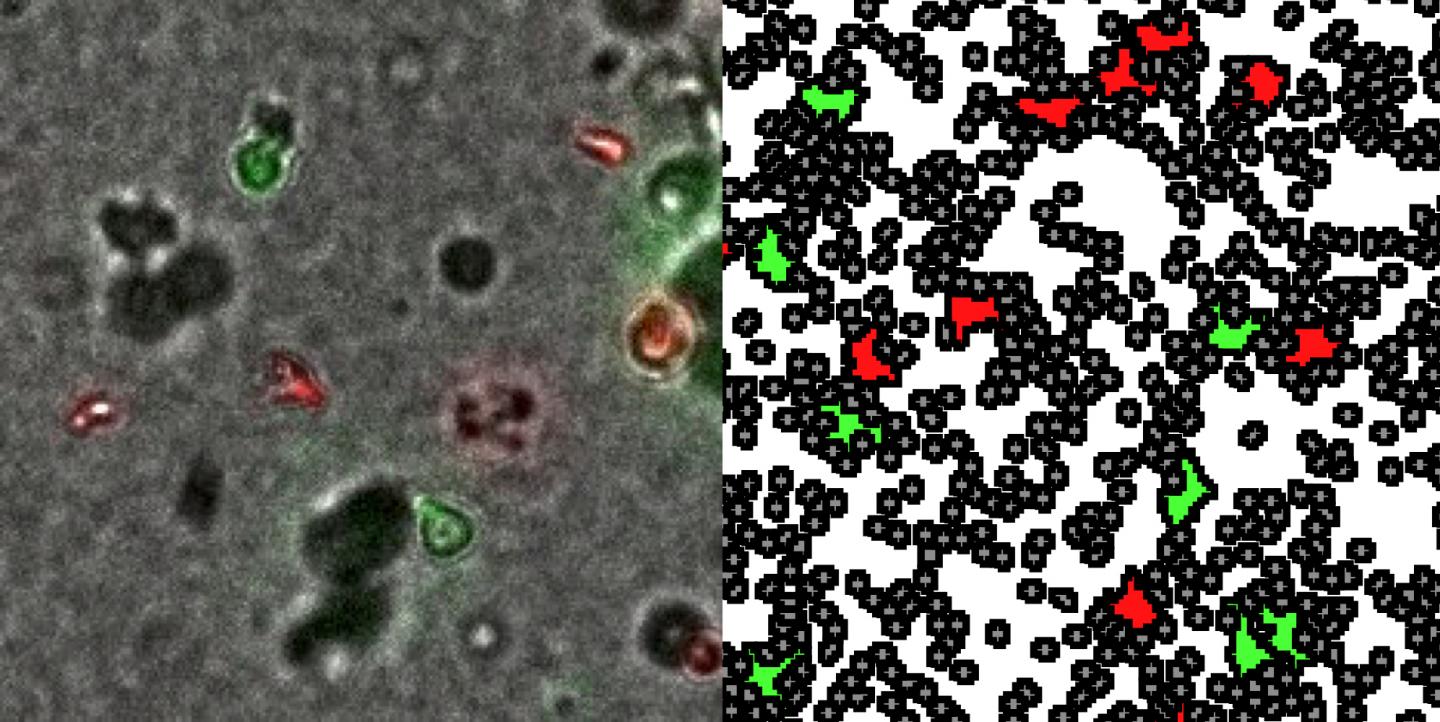
 {module In-article}
{module In-article}