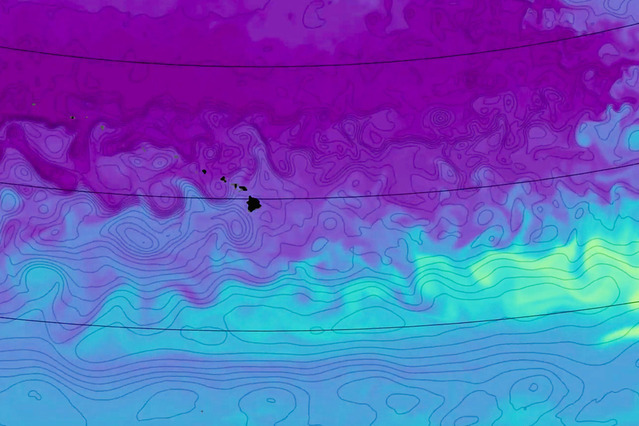
 Subtropical gyres are enormous rotating ocean currents that generate sustained circulations in the Earth’s subtropical regions just to the north and south of the equator. These gyres are slow-moving whirlpools that circulate within massive basins around the world, gathering up nutrients, organisms, and sometimes trash, as the currents rotate from coast to coast.
Subtropical gyres are enormous rotating ocean currents that generate sustained circulations in the Earth’s subtropical regions just to the north and south of the equator. These gyres are slow-moving whirlpools that circulate within massive basins around the world, gathering up nutrients, organisms, and sometimes trash, as the currents rotate from coast to coast.
For years, oceanographers have puzzled over conflicting observations within subtropical gyres. At the surface, these massive currents appear to host healthy populations of phytoplankton — microbes that feed the rest of the ocean food chain and are responsible for sucking up a significant portion of the atmosphere’s carbon dioxide.
But judging from what scientists know about the dynamics of gyres, they estimated the currents themselves wouldn’t be able to maintain enough nutrients to sustain the phytoplankton they were seeing. How, then, were the microbes able to thrive?
Now, MIT investigators have found that phytoplankton may receive deliveries of nutrients from outside the gyres and that the delivery vehicle is in the form of eddies — much smaller currents that swirl at the edges of a gyre. These eddies pull nutrients in from high-nutrient equatorial regions and push them into the center of a gyre, where the nutrients are then taken up by other currents and pumped to the surface to feed phytoplankton.
Ocean eddies, the team found, appear to be an important source of nutrients in subtropical gyres. Their replenishing effect, which the researchers call a “nutrient relay,” helps maintain populations of phytoplankton, which play a central role in the ocean’s ability to sequester carbon from the atmosphere. While climate models tend to project a decline in the ocean’s ability to sequester carbon over the coming decades, this “nutrient relay” could help sustain carbon storage over the subtropical oceans.
“There’s a lot of uncertainty about how the carbon cycle of the ocean will evolve as the climate continues to change, ” says Mukund Gupta, a postdoc at Caltech who led the study as a graduate student at MIT. “As our paper shows, getting the carbon distribution right is not straightforward, and depends on understanding the role of eddies and other fine-scale motions in the ocean.”
The study’s co-writers are Jonathan Lauderdale, Oliver Jahn, Christopher Hill, Stephanie Dutkiewicz, and Michael Follows at MIT, and Richard Williams at the University of Liverpool.
A snowy puzzle
A cross-section of an ocean gyre resembles a stack of nesting bowls that is stratified by density: Warmer, lighter layers lie at the surface, while colder, denser waters make up deeper layers. Phytoplankton lives within the ocean’s top sunlit layers, where the microbes require sunlight, warm temperatures, and nutrients to grow.
When phytoplankton dies, they sink through the ocean’s layers as “marine snow.” Some of this snow releases nutrients back into the current, where they are pumped back up to feed new microbes. The rest of the snow sinks out of the gyre, down to the deepest layers of the ocean. The deeper the snow sinks, the more difficult it is for it to be pumped back to the surface. The snow is then trapped, or sequestered, along with any unreleased carbon and nutrients.
Oceanographers thought that the main source of nutrients in subtropical gyres came from recirculating marine snow. But as a portion of this snow inevitably sinks to the bottom, there must be another source of nutrients to explain the healthy populations of phytoplankton at the surface. Exactly what that source is “has left the oceanography community a little puzzled for some time,” Gupta says.
Swirls at the edge
{media id=289,layout=solo}
In their new study, the team sought to simulate a subtropical gyre to see what other dynamics may be at work. They focused on the North Pacific gyre, one of the Earth’s five major gyres, which circulates over most of the North Pacific Ocean and spans more than 20 million square kilometers.
The team started with the MITgcm, a general circulation model that simulates the physical circulation patterns in the atmosphere and oceans. To reproduce the North Pacific gyre’s dynamics as realistically as possible, the team used a MITgcm algorithm, previously developed at NASA and MIT, which tunes the model to match actual observations of the ocean, such as ocean currents recorded by satellites, and temperature and salinity measurements taken by ships and drifters.
“We use a simulation of the physical ocean that is as realistic as we can get, given the machinery of the model and the available observations,” Lauderdale says.
The realistic model captured finer details, at a resolution of fewer than 20 kilometers per pixel, compared to other models that have a more limited resolution. The team combined the simulation of the ocean’s physical behavior with the Darwin model — a simulation of microbe communities such as phytoplankton, and how they grow and evolve with ocean conditions.
The team ran the combined simulation of the North Pacific gyre for over a decade, and created animations to visualize the pattern of currents and the nutrients they carried, in and around the gyre. What emerged were small eddies that ran along the edges of the enormous gyre and appeared to be rich in nutrients.
“We were picking up on little eddy motions, basically like weather systems in the ocean,” Lauderdale says. “These eddies were carrying packets of high-nutrient waters, from the equator, north into the center of the gyre and downwards along the sides of the bowls. We wondered if these eddy transfers made an important delivery mechanism.”
Surprisingly, the nutrients first move deeper, away from the sunlight, before being returned upwards where the phytoplankton live. The team found that ocean eddies could supply up to 50 percent of the nutrients in subtropical gyres.
“That is very significant,” Gupta says. “The vertical process that recycles nutrients from marine snow is only half the story. The other half is the replenishing effect of these eddies. As subtropical gyres contribute a significant part of the world’s oceans, we think this nutrient relay is of global importance.”
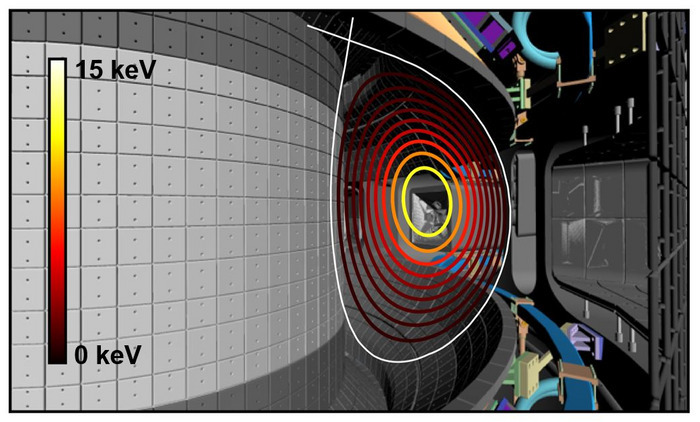 'FIRE mode' expected to resolve operational difficulties of commercial fusion reactors in the future
'FIRE mode' expected to resolve operational difficulties of commercial fusion reactors in the future

 How to resolve AdBlock issue?
How to resolve AdBlock issue? 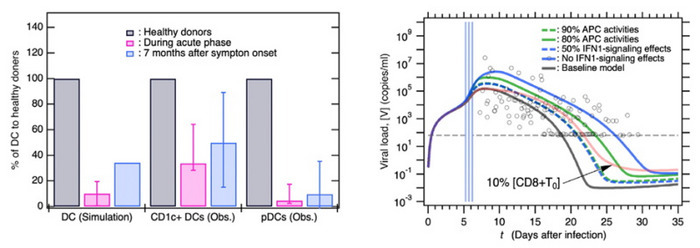 As COVID-19 wreaks havoc across the globe, one characteristic of the infection has not gone unnoticed. The disease is heterogeneous in nature with symptoms and severity of the condition spanning a wide range. The medical community now believes this is attributed to variations in the human hosts’ biology and has little to do with the virus per se. Shedding some light on this conundrum is Associate Professor SUMI Tomonari from Okayama University, Research Institute for Interdisciplinary Science (RIIS), and Associate Professor Kouji Harada from the Toyohashi University of Technology, the Center for IT-based Education (CITE). The duo recently reported their findings on imbalances in the host immune system that facilitate persistent or severe forms of the disease in some patients.
As COVID-19 wreaks havoc across the globe, one characteristic of the infection has not gone unnoticed. The disease is heterogeneous in nature with symptoms and severity of the condition spanning a wide range. The medical community now believes this is attributed to variations in the human hosts’ biology and has little to do with the virus per se. Shedding some light on this conundrum is Associate Professor SUMI Tomonari from Okayama University, Research Institute for Interdisciplinary Science (RIIS), and Associate Professor Kouji Harada from the Toyohashi University of Technology, the Center for IT-based Education (CITE). The duo recently reported their findings on imbalances in the host immune system that facilitate persistent or severe forms of the disease in some patients. Subtropical gyres are enormous rotating ocean currents that generate sustained circulations in the Earth’s subtropical regions just to the north and south of the equator. These gyres are slow-moving whirlpools that circulate within massive basins around the world, gathering up nutrients, organisms, and sometimes trash, as the currents rotate from coast to coast.
Subtropical gyres are enormous rotating ocean currents that generate sustained circulations in the Earth’s subtropical regions just to the north and south of the equator. These gyres are slow-moving whirlpools that circulate within massive basins around the world, gathering up nutrients, organisms, and sometimes trash, as the currents rotate from coast to coast.