A Bristol-led team of physicists has found a way to operate mass manufacturable photonic sensors at the quantum limit. This breakthrough paves the way for practical applications such as monitoring greenhouse gases and cancer detection.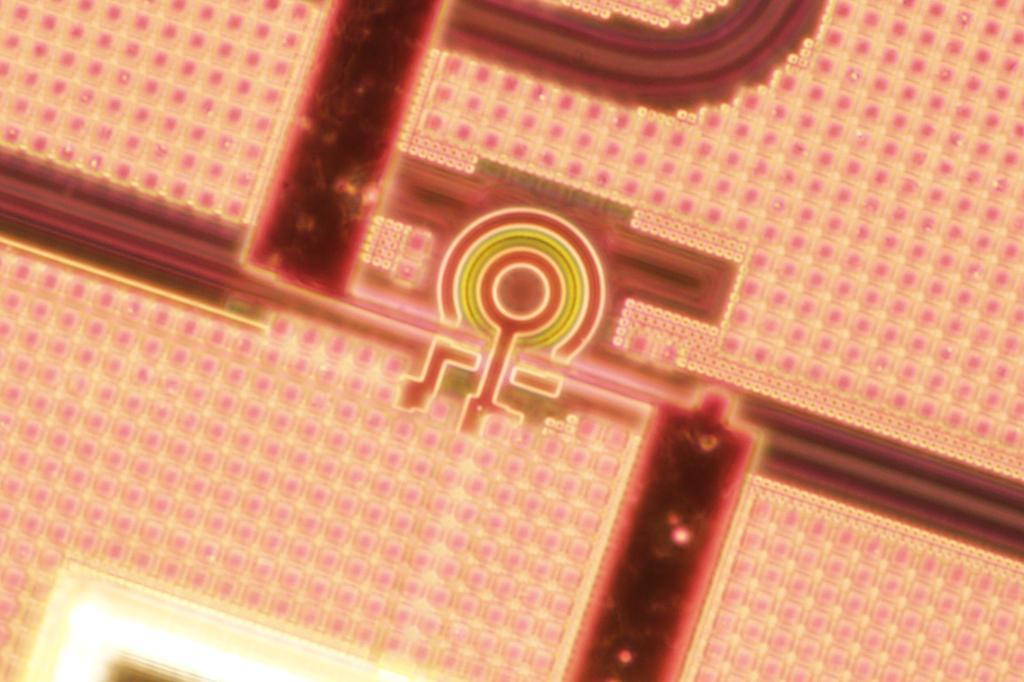 
Sensors are a constant feature of our everyday lives. Although they often go unperceived, sensors provide critical information essential to modern healthcare, security, and environmental monitoring. Modern cars alone contain over 100 sensors and this number will only increase.
Quantum sensing is poised to revolutionize today's sensors, significantly boosting the performance they can achieve. More precise, faster, and reliable measurements of physical quantities can have a transformative effect on every area of science and technology, including our daily lives.
However, the majority of quantum sensing schemes rely on special entangled or squeezed states of light or matter that are hard to generate and detect. This is a major obstacle to harnessing the full power of quantum-limited sensors and deploying them in real-world scenarios.
In a paper published today, a team of physicists at the Universities of Bristol, Bath, and Warwick have shown it is possible to perform high precision measurements of important physical properties without the need for sophisticated quantum states of light and detection schemes.
The key to this breakthrough is the use of ring resonators – tiny racetrack structures that guide light in a loop and maximize its interaction with the sample under study. Importantly, ring resonators can be mass-manufactured using the same processes as the chips in our computers and smartphones.
Alex Belsley, Quantum Engineering Technology Labs (QET Labs) Ph.D. student and lead author of the work, said: “We are one step closer to all integrated photonic sensors operating at the limits of detection imposed by quantum mechanics.”
Employing this technology to sense absorption or refractive index changes can be used to identify and characterize a wide range of materials and biochemical samples, with topical applications from monitoring greenhouse gases to cancer detection.
Associate Professor Jonathan Matthews, co-Director of QET Labs and co-author of the work, stated: “We are really excited by the opportunities this result enables: we now know how to use mass manufacturable processes to engineer chip-scale photonic sensors that operate at the quantum limit.”


 How to resolve AdBlock issue?
How to resolve AdBlock issue?