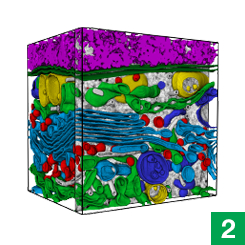
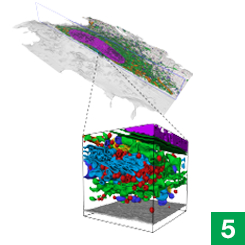
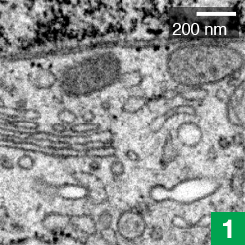
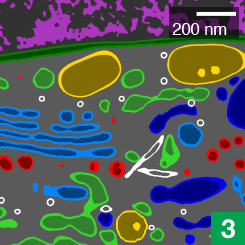
Janelia’s COSEM Project Team has created a set of tools to make annotated 3D images of cells, showing the relationships between different organelles.
Open any introductory biology textbook, and you’ll see a familiar diagram: A blobby-looking cell filled with brightly colored structures – the inner machinery that makes the cell tick.
Cell biologists have known the basic functions of most of these structures, called organelles, for decades. The bean-shaped mitochondria make energy, for example, and lanky microtubules help cargo zip around the cell. But for all that scientists have learned about these miniature ecosystems, much remains unknown about how their parts all work together. 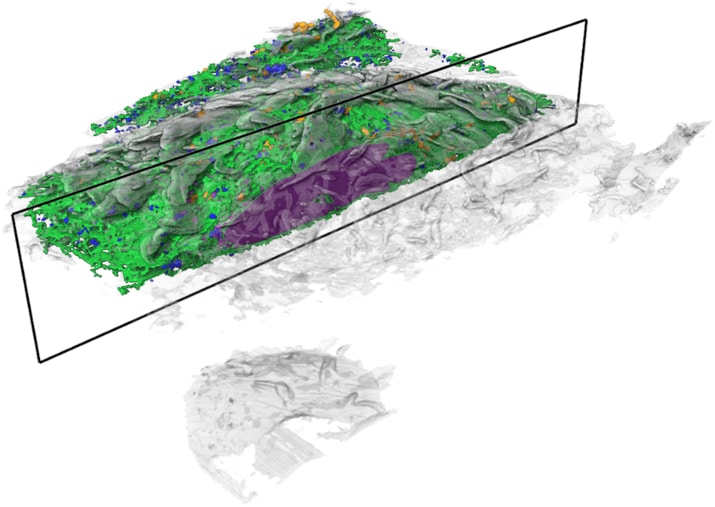
Now, high-powered microscopy – plus a heavy dose of machine learning – is helping to change that. New computer algorithms can automatically identify some 30 different kinds of organelles and other structures in super-high-resolution images of entire cells, a team of scientists at the Howard Hughes Medical Institute’s Janelia Research Campus reports.
The detail in these images would be nearly impossible to parse by hand throughout the entire cell, says Aubrey Weigel, who led the Janelia Project Team, called COSEM (for Cell Organelle Segmentation in Electron Microscopy). The data for just one cell is made up of tens of thousands of images; tracing all a cell’s organelles through that collection of pictures would take one person more than 60 years. But the new algorithms make it possible to map an entire cell in hours, rather than years.
{media load=media,id=258,width=350,align=left,display=inline}
“By using machine learning to process the data, we felt we could revisit the canonical view of a cell,” Weigel says.
Janelia scientists also released a data portal, OpenOrganelle, where anyone can access the datasets and tools they’ve created.
These resources are invaluable for scientists studying how organelles keep cells running, says Jennifer Lippincott-Schwartz, a senior group leader and interim head of Janelia’s new 4D Cellular Physiology research area who is already using the data in her research. “What we haven’t really known is how different organelles and structures are arranged relative to each other – how they’re touching and contacting each other, how much space they occupy,” she says.
For the first time, those hidden relationships are visible.
Detailed data
The COSEM team’s journey started with data collected by high-powered electron microscopes housed in a special vibration-proof room at Janelia.
For the past ten years, these microscopes have been churning out high-resolution snapshots of the fly brain. Janelia Senior scientist Shan Xu and senior group leader Harald Hess have engineered these scopes to mill off super-thin slivers of the fly brain using a focused beam of ions – an approach called FIB-SEM imaging. The scopes capture images layer by layer, and then computer programs stitch those images together into a detailed 3D representation of the brain. Based on these data, Janelia researchers released the most detailed neural map of the fly brain yet.
While imaging the fly brain, Hess and Xu’s team also looked at other samples. Over time, they amassed a collection of data from many kinds of cells, including mammalian cells. “We thought that this detailed imaging of whole cells might be of larger interest to cell biologists,” Hess says.
{media load=media,id=259,width=350,align=left,display=inline}
Weigel, then a postdoc in Lippincott-Schwartz’s lab, began mining those data for her research. “The resolving power of the FIB-SEM imaging was amazing, and we were able to see things at a level we couldn’t have imagined before,” says Weigel, “but there was more information in one sample than I could analyze in several lifetimes.” Realizing that others at Janelia were working on computational projects that might speed things up, she began organizing a collaboration.
“All of the pieces were here at Janelia,” she says, and forming the COSEM project team aligned them towards a common goal.
Setting boundaries
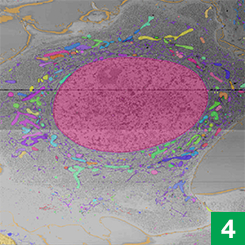
Larissa Heinrich, a graduate student in group leader Stephan Saalfeld’s lab, had previously developed machine learning tools that could pinpoint synapses, the connections between neurons, in electron microscope data. For COSEM, she adapted those algorithms to instead map out, or segment, organelles in cells.
Saalfeld and Heinrich’s segmentation algorithms worked by assigning each pixel in an image a number. The number reflected how far the pixel was from the nearest synapse. An algorithm then used those numbers to ID and label all the synapses in an image. The COSEM algorithms work similarly, but with more dimensions, Saalfeld says. They classify every pixel by its distance to each of 30 different kinds of organelles and structures. Then, the algorithms integrate all of those numbers to predict where organelles are positioned.
Using data from scientists who have manually traced organelle boundaries and assigned numbers to pixels, the algorithm can learn that particular combinations of numbers are unreasonable, Saalfeld says. “So, for example, a pixel can’t be inside a mitochondrion at the same time it’s inside the endoplasmic reticulum.”
To answer questions like how many mitochondria are in a cell, or what their surface area is, the algorithms need to go even further, says group leader Jan Funke. His team built algorithms that incorporate prior knowledge about organelles’ characteristics. For example, scientists know that microtubules are long and thin. Based on that information, the computer can make judgments about where a microtubule begins and ends. The team can observe how such prior knowledge affects the computer program’s results – whether it makes the algorithm more or less accurate – and then make adjustments where necessary.
After two years of work, the COSEM team has landed on a set of algorithms that generate good results for the data that have been collected so far. Those results are the important groundwork for future research at Janelia, says Weigel. A new effort headed by Xu is taking FIB-SEM imaging to even greater levels of detail. And another soon-to-launch project team named CellMap will further refine COSEM’s tools and resources to create a more expansive cell annotation database, with detailed images of many more types of cells and tissue.
Together, those advances will support Janelia’s next 15-year research area, 4D Cellular Physiology – an effort led on an interim basis by Lippincott-Schwartz to understand how cells interact with each other within each of the many different kinds of tissue that make up an organism, says Wyatt Korff, Director of Project Teams at Janelia.
With new resources like those created by the COSEM team and the Enhanced FIB-SEM Technology group, Korff says, “we can actually begin to answer those questions, in a way that we haven’t had access to in the past.”



 How to resolve AdBlock issue?
How to resolve AdBlock issue?