An evocative new image captured by the Gemini North telescope in Hawai‘i reveals a pair of interacting spiral galaxies — NGC 4568 and NGC 4567 — as they begin to clash and merge. These galaxies are entangled by their mutual gravitational field and will eventually combine to form a single elliptical galaxy in around 500 million years. Also visible in the image are the glowing remains of a supernova that was detected in 2020.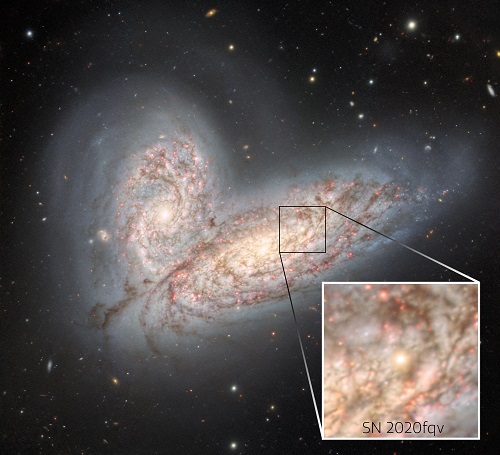 
Gemini North, one of the twin telescopes of the International Gemini Observatory, operated by NSF’s NOIRLab, has observed the initial stages of a cosmic collision approximately 60 million light-years away in the direction of the constellation Virgo. The two stately spiral galaxies, NGC 4568 (bottom) and NGC 4567 (top) are poised to undergo one of the most spectacular events in the Universe, a galactic merger. At present, the centers of these galaxies are still 20,000 light-years apart (about the distance from Earth to the center of the Milky Way) and each galaxy still retains its original, pinwheel shape. Those placid conditions, however, will change.
As NGC 4568 and NGC 4567 draw together and coalesce, their dueling gravitational forces will trigger bursts of intense star formation and wildly distort their once-majestic structures. Over millions of years, the galaxies will repeatedly swing past each other in ever-tightening loops, drawing out long streamers of stars and gas until their structures are so thoroughly mixed that a single, essentially spherical, galaxy emerges from the chaos. By that point, much of the gas and dust (the fuel for star formation) in this system will have been used up or blown away.
This merger is also a preview of what will happen when the Milky Way and its closest large galactic neighbor the Andromeda Galaxy collide in about 5 billion years.
A bright region in the center of one of NGC 4568’s sweeping spiral arms is the fading afterglow of a supernova — known as SN 2020fqv — that was detected in 2020. The new Gemini image was produced from data taken in 2020.
By combining decades of observations and supercomputer modeling, astronomers now have compelling evidence that merging spiral galaxies like these go on to become elliptical galaxies. Likely, NGC 4568 and NGC 4567 will eventually resemble their more-mature neighbor Messier 89, an elliptical galaxy that also resides in the Virgo Cluster. With its dearth of star-forming gas, Messier 89 now exhibits minimal star formation and is made up primarily of older, low-mass stars and ancient globular clusters.
Advanced technology on the Gemini North telescope, including the Gemini Multi-Object Spectrograph North (GMOS-N) and the dry air above the summit of Maunakea, allowed astronomers to capture this spectacular image.
The image was obtained by NOIRLab’s Communication, Education & Engagement team, as part of the NOIRLab Legacy Imaging Program.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?