Breaks the limitations of classical mechanics by introducing a quasi-quantum effect; expected to accelerate complex problem-solving in finance, pharmaceuticals, and logistics
Toshiba Corporation (TOKYO: 6502) and Toshiba Digital Solutions Corporation (collectively Toshiba), industry leaders in solutions for large-scale optimization problems, today announced the Ballistic Simulated Bifurcation Algorithm (bSB) and the Discrete Simulated Bifurcation Algorithm (dSB), new algorithms that far surpass the performance of Toshiba's previous Simulated Bifurcation Algorithm (SB). The new algorithms will be applied to finding solutions to highly complex problems in areas as diverse as portfolio management, drug development and logistics management.
Introduced in April 2019, the previous SB broke new ground as a platform for finding solutions to combinatorial optimization problems, surpassing other approaches by a factor of 10*1. Toshiba has now extended this achievement with two new algorithms that apply innovative approaches, such as a quasi-quantum tunneling effect, to performance improvement, allowing them to acquire optimal solutions (exact solutions) for large-scale combinatorial optimization problems that challenge the capabilities of their predecessor. Implemented on a 16-GPU machine, dSB can find a nearly optimal solution of a one-million-bit problem, the world's largest scale combinatorial problem yet reported in scientific papers, in 30 minutes--a computation that would take 14 months on a typical CPU-based computer. The research results were published in the online academic journal, Science Advances.
The new algorithms have different characteristics. bSB is optimized and named for speed of operation and finds good approximate solutions in a short time. It generates fewer errors than a previously reported Adiabatic Simulated Bifurcation Algorithm (aSB)*3, and so returns faster, more accurate results. Implemented on a field-programmable gate array (FPGA), dubbed the ballistic simulated bifurcation machine (bSBM), it obtains a good solution to a 2,000-bit problem approximately 10 times faster than the previous aSB machine (aSBM) (Figure 1). 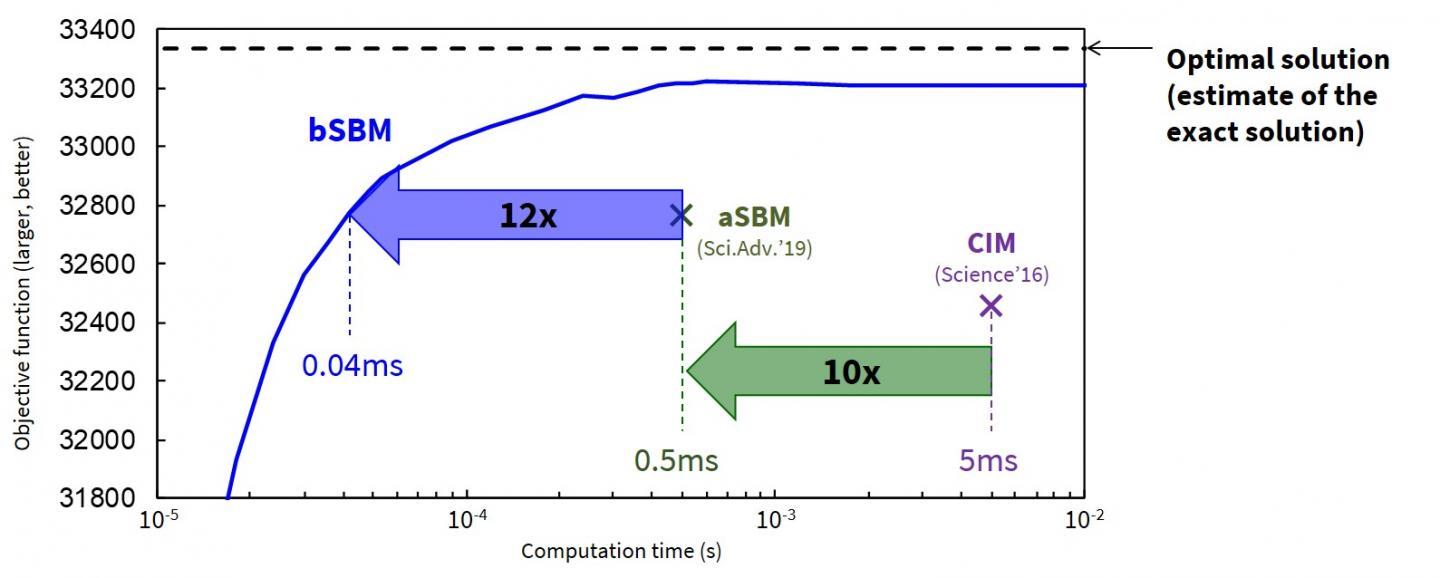 {module INSIDE STORY}
{module INSIDE STORY}
dSB is a high-accuracy algorithm. Although implemented in a classical computer, it nonetheless arrives at optimal solutions faster than current quantum machines. Its name is derived from the replacement of continuous variables with discrete variables in equations of motion. This exhibits a quasi-quantum tunneling effect that breaks through the limits of approaches grounded in classical mechanics, reaching the optimal solution of the 2000-bit problem.
Toshiba has implemented dSB on a FPGA and built a discrete simulated bifurcation machine (dSBM) that achieves a higher speed than other machines in terms of computation times required to obtain optimal solutions for various problems (Figure 2). 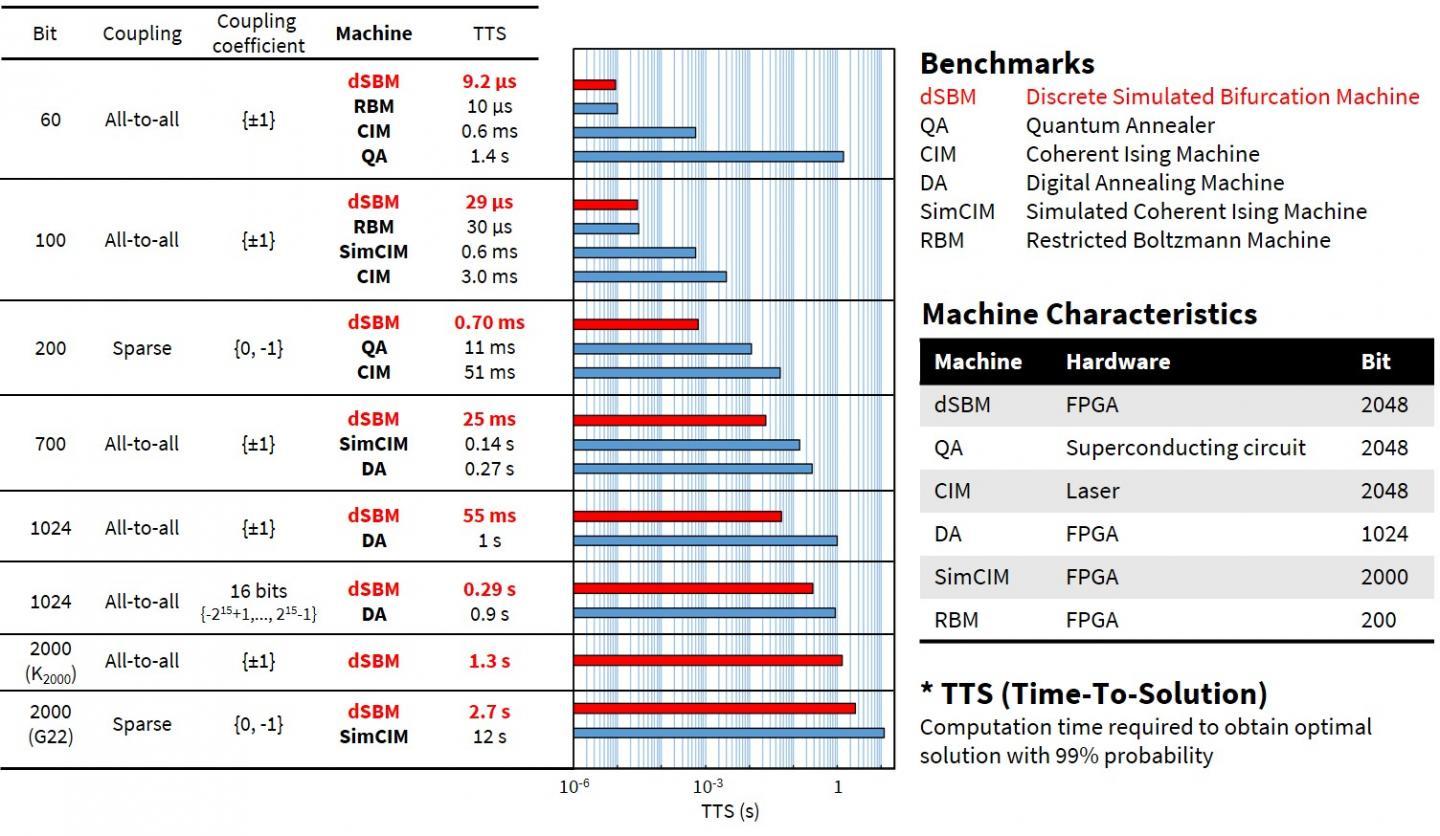 {module INSIDE STORY}
{module INSIDE STORY}
Implemented on a 16-GPU machine, the dSBM solved a one-million-bit problem, the largest yet reported in scientific papers, and arrived at a nearly optimal solution in 30 minutes--20,000 times faster than a CPU-based simulated annealing machine, which would take 14 months to carry out the computation (Figure 3). 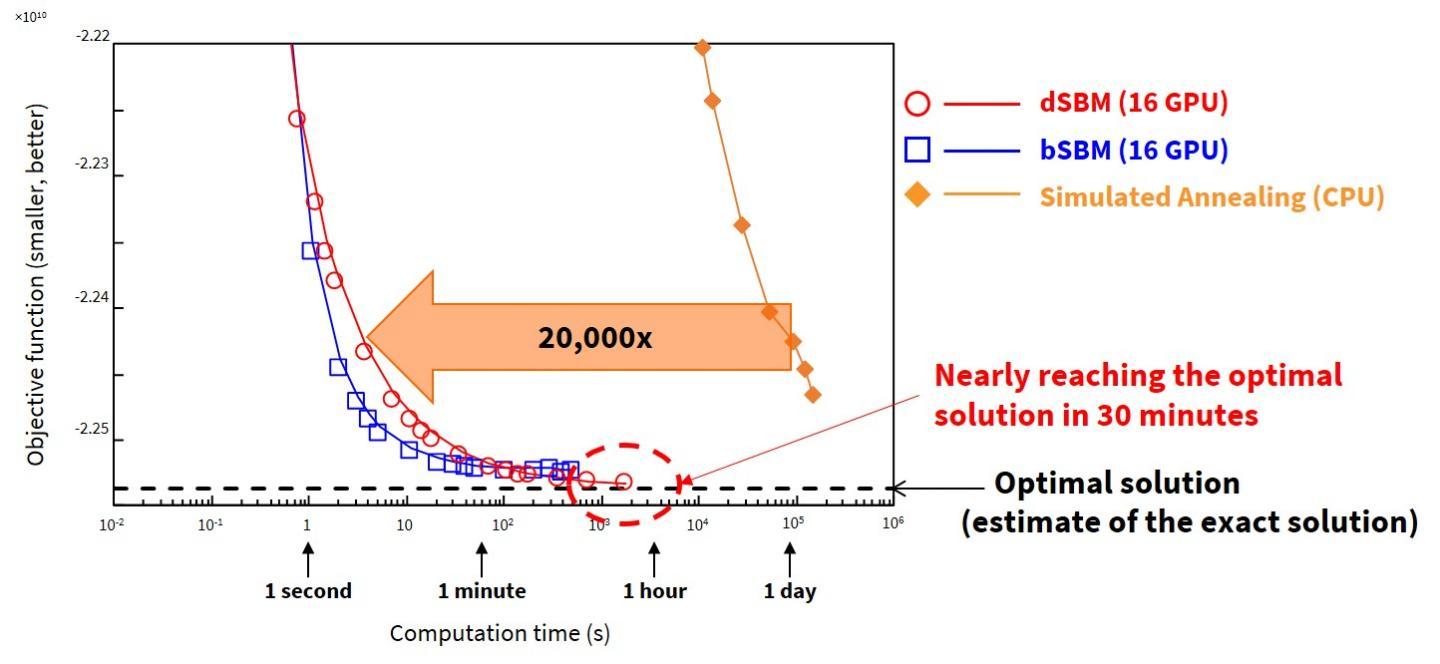
In applying the two algorithms to real-world problems, Toshiba proposes bSB for applications that require an immediate response, and dSB for applications that require high accuracy, even if it takes a little longer time.
Toshiba expects the new algorithms to bring higher efficiencies to industry, business and complex decision-making by addressing combinatorial optimization problems in fields including investment portfolios, drug development, and delivery route planning.
Commenting on the algorithms, Hayato Goto, Chief Research Scientist at Toshiba Corporation's Corporate Research & Development Center, said: "We face many real-world problems where we must find the optimal solution among a huge number of choices, and we must also deal with combinatorial explosion, where the number of combination patterns increases exponentially as a problem increases in scale. This is why research into special-purpose computers for combinatorial optimization is being carried out worldwide. Our aim is to develop a software solution--algorithms that can solve large-scale combinatorial optimization problems quickly and accurately, and contribute to the realization of higher efficiencies."
Toshiba will offer the newly developed simulated bifurcation algorithms as a GPU-based cloud service and as an on-premises version implemented on an FPGA within 2021.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module INSIDE STORY}
{module INSIDE STORY}