According to a new model, if the U.S. presidential election were to take place today, former Vice President Joe Biden would have an 88.3% percent chance of winning. That’s the finding of a group of U.S. university researchers based on new research published in SIAM Review today.
This finding assumes that Americans vote the way that they say they will in publicly available polling data and that voters not accounted for in existing polling data will turn out equally for both candidates.
How did the researchers — from Northwestern University, UCLA, Augusta University, and The Ohio State University — arrive at their conclusion? By applying a modeling framework like one's experts use to forecast the spread of infectious diseases (such as COVID-19) to the high-stakes challenge of forecasting election outcomes.
“What we assume is that similar to how an infected person can cause — or influence — a susceptible person to become infected with a virus, a Republican or Democratic voter can influence an undecided voter,” said lead researcher Alexandria Volkening, an NSF–Simons Fellow at Northwestern University, who co-authored the study with UCLA Mathematics professor Mason Porter, Augusta University Biostatistics and Data Science professor Daniel Linder, and Ohio State University Biostatistics and Mathematics professor Grzegorz Rempala. Their 2020 forecasts are also done in collaboration with Volkening’s students Samuel Chian, William He, and Christopher Lee.
“I think we were all initially surprised that a disease-transmission model could produce meaningful forecasts of elections, but one of the benefits of mathematical modeling is that you can apply similar methods to shed light on many different problems,” she added.
The group’s election forecasting model — which is based on “compartmental modeling” — was shown to have a similar success rate to popular forecasters FiveThirtyEight and Sabato’s Crystal Ball.
Researchers treated Democratic and Republican voting inclinations as two possible kinds of ‘infections’ that can spread between states. Undecided, independent, or minor-party voters were considered ‘susceptible’ individuals, and infection was interpreted as adopting Democratic or Republican opinions. ‘Recovery’ represented the turnover of committed voters to undecided ones.
Unlike election forecasts that combine polling data with other data, such as historical voting, the economy, and approval ratings, the researchers’ model uses only publicly available polling data and treats all polls on equal footing. Transmission is interpreted as opinion persuasion, influenced by campaigning, media coverage and debates, and opinions spread both within and between states. Despite its simplicity, the model performs surprisingly well, Volkening explained. For example, it was as effective as popular analysts were at predicting (known as “calling”) the 2012 and 2016 races for governors, senators, and presidents in the U.S. using historical polling data, she said.
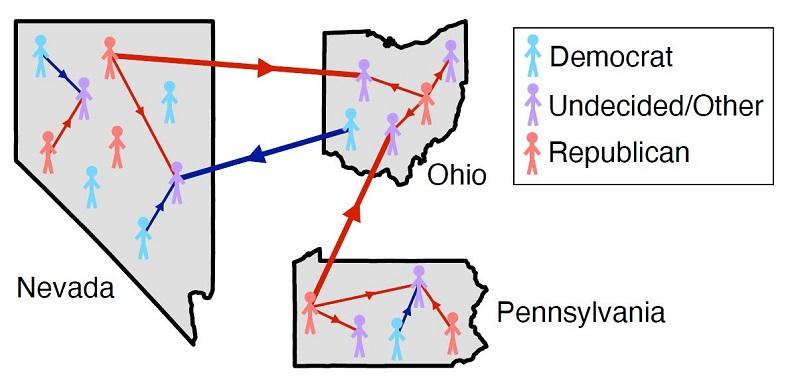
Figure 1: Voters can interact both within and between states, influencing each other’s political opinions. Figure courtesy of Alexandria Volkening, Daniel F. Linder, Mason A. Porter, and Grzegorz A. Rempala. {module INSIDE STORY}
"One important limitation is that we assume all undecided individuals who are left at the end of our simulated elections vote for minor-party candidates or turn out equally for the Democratic and Republican candidates,” Volkening said. “If undecided voters all vote in one direction or voter turnout is heavily partisan, it is very possible for a trailing candidate to win.”
Though the paper is being published in the midst of a global pandemic, UCLA’s Porter is quick to point out that the idea to use a disease transmission model was made long before the COVID-19 pandemic surfaced.
“When we first discussed using this approach, it was on the heels of the 2016 election when pollsters were predicting a Clinton win and of course that’s not what happened,” said Porter, noting that the researchers speculated that something was wrong with the forecasting models that were being applied, the polling data itself, or the interpretations of forecast uncertainty.
“There are many tools already available for compartmental modeling because people have studied infectious diseases for quite some time with great success, so it made sense to try a similar approach to study election forecasting,” he said.
The group’s model and U.S. election forecasts are publicly available at https://modelingelectiondynamics.gitlab.io/2020-forecasts/index.html and the researchers strongly encourage readers to try out their modeling framework and build on it further.
To read the entire study, visit SIAM Review.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module INSIDE STORY}
{module INSIDE STORY}